In a new study, researchers from research institutions such as the Walter Reed Army Research Institute in the United States confirmed previous research on host limiting factors for HIV-1. They detailed how omics reveal correlations when searching for treatment strategies. The relevant research results were published in the issue of the Science Translational Medicine journal, with the title “Single-cell transcriptomics identifies prothymosin α restriction of HIV-1 in vivo.”
These authors used single-cell transcriptomics to identify proteins active in the cells of 14 HIV-1 infected individuals. They narrowed down the list of potential antiviral proteins called host restriction factors by correlating the patient’s gene expression with the number of HIV-1 RNA in a single cell. These proteins are part of the innate immune system in the human body and can recognize and interfere with specific steps in the virus replication cycle, thereby preventing infection.
These authors also sequenced and compared one participant with the highest plasma viral load, and found that the transcription of HIV-1 RNA is inversely proportional with the expression of ProTα (prothymosin αlpha), encoding by the gene PTMA. This association was subsequently validated among 28 participants in addition to this initial study. It has been confirmed that overexpression of ProTα in vitro, this cytokine can inhibit the transcription of HIV-1 and the production of infectious viruses.
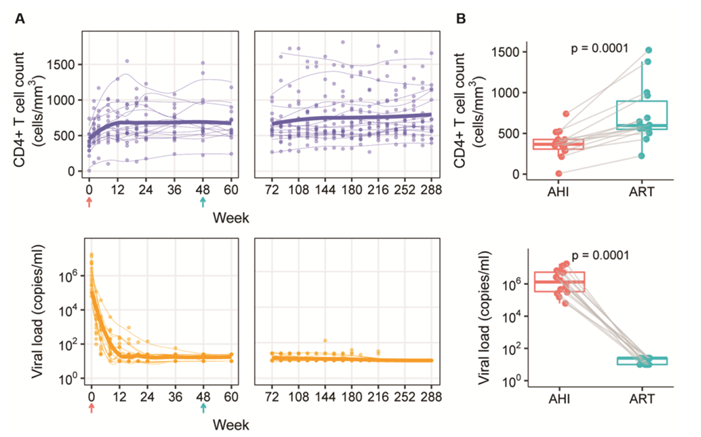
These authors found that HIV-1 RNA in individual immune cells is associated with specific clinical measurement results. Through single-cell analysis, they identified cells carrying HIV-1 RNA from participants during acute HIV infection (AHI) and receiving antiretroviral therapy (ART).
HIV-1 RNA-positive cells are mostly present in the CD4+T cell subpopulation. Correlation analysis shows that the frequency of HIV-1 RNA-positive CD4+T cells is positively correlated with indicators such as the total amount of cell-related HIV-1 DNA and plasma viral load. This indicates that the presence of HIV-1 RNA in these cells has biological significance and is related to HIV-1 clinical parameters.
In confirming this method, researchers at Mount Sinai Medical School in New York discovered the limitations of ProTα on HIV-1 infection 17 years ago, which were published in the Journal of Virology titled “Novel Function of Prothymosin Alpha as a Potent Inhibitor of Human Immunodeficiency Virus Type 1 Gene Expression in Primary Macrophages”. They also observed that after human T cells were infected with HIV-1, the detectable measurement of ProTα mRNA is reduced. They discovered that the active ProTα significantly inhibited HIV-1 replication in primary macrophages in a dose-dependent manner, but had almost no effect on HIV-1 in CD4+T cells.
This new study confirms that unbiased omics techniques can be used to identify potential antiviral factors, such as ProTα targeting HIV-1, and other virus prevention and treatment strategies.
Related Services
Protein Expression and Purification Services
Reference
Aviva Geretz et al. Single-cell transcriptomics identifies prothymosin α restriction of HIV-1 in vivo. Science Translational Medicine, 2023, doi:10.1126/scitranslmed.adg0873.