Proteomic Analysis Kits
🧪 YNI-016K
Source:
Species:
Tag:
Conjugation:
Protein Length:
Background
Overview
Proteomics is an important branch of life science, which involves the comprehensive analysis of all proteins in living organisms, including protein expression, structure, function and interaction. Specifically, the contents of proteomics research mainly include the following aspects:
Protein identification: The identification of all proteins present in a cell or tissue, including their types and quantities.
Protein quantification: Measure protein expression levels and analyze changes in protein abundance under different states.
Protein localization: Study the distribution of proteins within cells to understand their subcellular localization.
Post-translational modification: Analysis of the chemical modifications of proteins after translation, such as phosphorylation, ubiquitination, methylation, and acetylation, which have important effects on the activity, stability, and function of proteins.
Functional proteomics: The study of the biological function of specific proteins or protein networks.
Structural proteomics: The study of the three-dimensional structure of proteins through structural biology methods to provide a basis for understanding their function.
Protein-protein interactions: Study of the interaction network between proteins, which is essential for understanding biological processes such as signal transduction and cell cycle regulation.
Disease proteomics: The search for disease markers and potential therapeutic targets by comparing proteomic differences between disease and normal states.
Drug discovery and development: Use proteomics data to discover new drug targets and develop drugs that target specific proteins.
Biomarker discovery: Identification of protein markers for disease diagnosis and prognostic assessment.
Systems biology: Conducting system-wide investigations to identify potential biomarkers and therapeutic targets for disease pathways and host-pathogen interactions.
Kits Types
A variety of detection kits are used in proteomics research, which are mainly used for protein extraction, quantification, separation, enrichment, detection and analysis. Here are some common proteomic test kits and their uses:
Protein extraction kits: Used to extract total protein from cell or tissue samples, such as by using SDS, NP40, TritonX100, Urea and other detergent to destroy cell structure and release protein.
Protein quantification kits: Such as BCA or Bradford protein quantification kits, are used to determine the concentration of protein solutions for subsequent enzymatic reactions and mass spectrometry.
Proteolytic kits: Provides enzymolytic reagents such as trypsin for cutting proteins into peptide segments in preparation for mass spectrometry.
Phosphorylated protein enrichment kits: Used to enrich phosphorylated modified peptides, these kits usually contain specific antibodies to recognize and enrich modified peptides.
HPLC (High performance liquid chromatography) separation kits: Used for fractional separation of peptide segments to improve the accuracy and depth of mass spectrometry detection.
Signal molecule kits: Used to screen and quantify multi-component hundreds of thousands of protein sites at key signal transduction nodes, critical for drug discovery, biomarker discovery and diagnostic method development.
Proteomic mass spectrometry Kits: For clinical protein mass spectrometry, which may include specific sample pretreatment and labeling techniques.
Importances of Proteomics
Proteomics helps scientists understand the complex networks and interactions of proteins within cells, revealing the molecular mechanisms involved in life processes. By comparing differences in protein expression between healthy and diseased states, proteomics helps to discover disease-related biomarkers and potential therapeutic targets. Proteomics plays a key role in drug design and screening and can be used to identify molecular targets of drug action and evaluate the effects and side effects of drugs. Proteomics technology helps to develop new diagnostic methods, improve the accuracy and efficiency of disease diagnosis, and provide molecular basis for personalized medicine. Proteomics studies can discover and validate disease-specific biomarkers that can be used for early disease diagnosis, prognostic assessment, and therapeutic response monitoring.
In agriculture, proteomics helps to improve crops, improve disease resistance and nutritional value of crops, and is also used for food quality control and safety testing. In environmental science, proteomics can be used to assess the effects of environmental factors on organisms, such as the effects of pollutants on protein expression in aquatic organisms. The analysis of proteomic data drives the development of bioinformatics and computational biology, facilitating the innovation of algorithms and computational models.
Proteomics contributes to the study of brain function and neurodegenerative diseases such as Alzheimer's and Parkinson's, advancing the development of neuroscience. In tumor immunology, proteomics contributes to understanding how the immune system recognizes and responds to tumors, informing the development of new cancer immunotherapies. Proteomics is an important part of systems biology research and contributes to a comprehensive understanding of the complexity and dynamics of biological systems. Proteomics can be used in forensic science for individual identification and toxicology studies, analyzing the effects of poisons or drugs on living organisms.

Applications
Protein extraction: Using specific lysis buffers and kits, cells can be efficiently lysed and proteins extracted.
Protein quantification: The protein concentration is quantified by a kit such as BCA or Bradford to ensure consistency and comparability of protein mass in subsequent experiments.
Protein enzymolysis: The use of enzymolysis kits such as trypsin to cut proteins into peptide segments in preparation for mass spectrometry.
Post-translational modification (PTM) enrichment: Enrichment of specific PTMS, such as peptides modified by phosphorylation, ubiquitination, and glycosylation, using kits that typically contain specific antibodies or ligands for affinity purification.
Single-cell proteomics: Mass spectroscope-based single-cell proteomics techniques enable researchers to make the transition from large sample analysis to single-cell analysis, contributing to a deeper understanding of cellular heterogeneity.
Viral studies: Mass spectrometry studies exploring the complete viral particle, surface, binding, protein composition, and their effects on host cell biochemical pathways.
High-throughput proteomic analysis: Illumina has partnered with SomaLogic to develop multiple high-throughput proteomic analysis methods for NGS instrument reading, further expanding the scope of high-throughput proteomic applications.

Case Study
Case Study 1: Total Protein Assay Kit (Kit-0731)
Nephrotic syndrome (NS) is one of the most commonly encountered glomerular diseases in children and a major contributor to the workload of pediatric nephrologists. Nephrotic syndrome is a chronic disease that is well known for its long term serious complications. among these complications albuminemia and dyslipidemia. Many studies has related albuminemia and dyslipidemia to other complications of Nephrotic syndrome as retinopathy and cardiovascular disease of nephrotic syndrome. The main objective of this study was planned to elucidate the potential diagnostic role of transforming growth factor-beta (TGF-β) as a novel biomarkers and its correlation with albuminemia and dyslipidemia in nephrotic syndrome among Egyptian children. Serum level of TGF-β was measured using ELISA, serum level of lipid profile and urinary proteins were estimated. This study revealed that serum level of TGF-β was higher in nephrotic cases than controls and its serum level was significantly negative correlated with serum albumin while it showed a significant positive correlation with urinary protein and lipid indices levels in children with nephrotic syndrome.
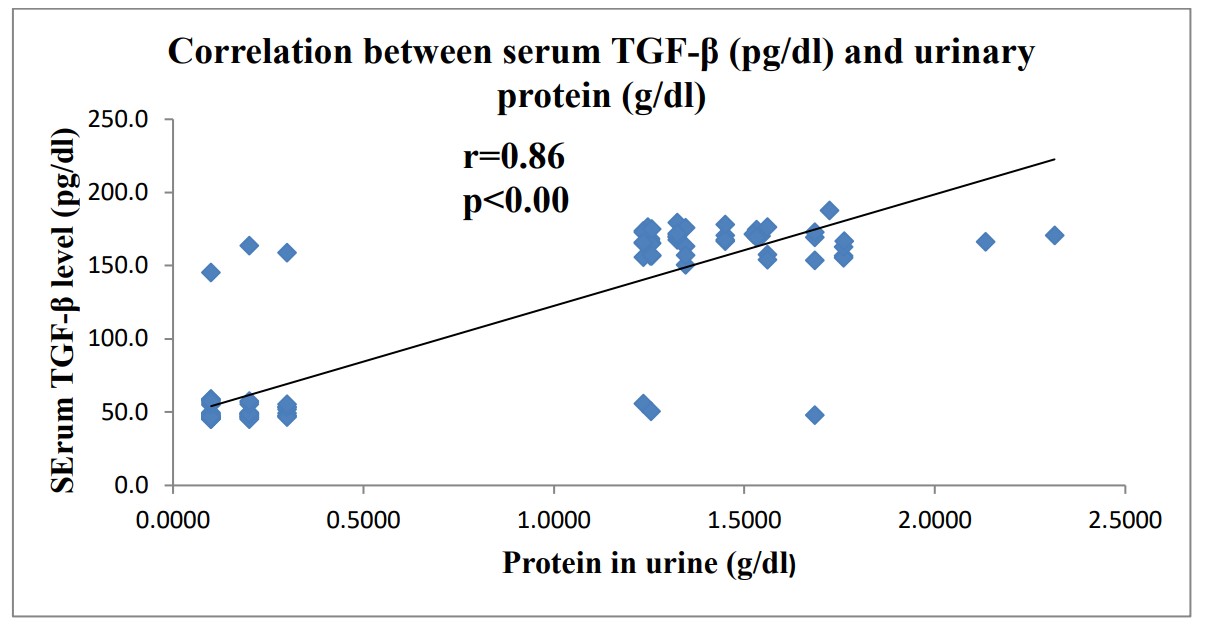
Fig1. Correlation between Serum TGF-β and Urinary Total Protein. (Faten Z. Mohamed, 2019)
Case Study 2: BCA Protein Assay Kit (Kit-0731)
Tear sampling is an attractive option for collecting biological samples in ophthalmology clinics, as it offers a non-invasive alternative to other invasive techniques. However, there are many tear sampling methods still in consideration. This study explores the suitability of Schirmer's test strip and microcapillary tube as reliable and satisfactory methods for tear sampling. The total tear protein concentrations were analyzed via spectrophotometry and bicinchoninic acid (BCA) protein assay. The protein profile was determined by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE). The optimal wetting length of Schirmer's strip and suitable buffer solutions were compared. The mean total protein obtained from an optimum wetting length of 20 mm using Schirmer's strip without anesthesia in phosphate-buffered saline (PBS) yielded substantial quantities of protein as measured by nanophotometer (220.20 ± 67.43 µg) and the BCA protein assay (210.34 ± 59.46 µg). This method collected a significantly higher quantity of protein compared to the microcapillary tube method (p=0.004) which was much more difficult to standardize. Using anaesthesia did not ease the sampling procedure substantially and protein quantity was maintained.
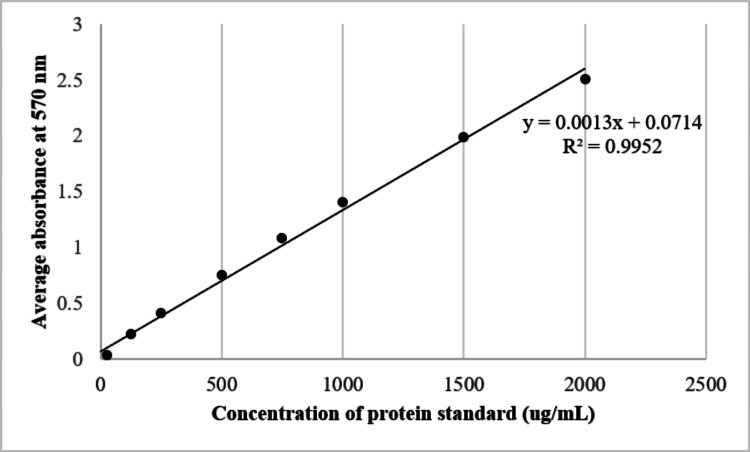
Fig2. The dispersion of the standard curve and its equation for average absorbance obtained at 570 nm using BCA Protein Assay Kit for BSA. (May Ling Tham, 2023)
Case Study 3: Caspase-3 Assay Kit
It has been reported that ginsenoside Rg1 (G‑Rg1) can alleviate alcoholic liver injury, cardiac hypertrophy and myocardial ischemia, as well as reperfusion injury. Therefore, the present study aimed to investigate the role of G‑Rg1 in alcohol‑induced myocardial injury, as well as to elucidate its underlying mechanisms of action. For this purpose, H9c2 cells were stimulated with ethanol. Subsequently, the levels of lactate dehydrogenase and caspase‑3 in the H9c2 cell culture supernatant were detected using corresponding assay kits. In addition, the expression of green fluorescent protein (GFP)‑light chain 3 (LC3) and that of C/EBP homologous protein (CHOP) were evaluated using GFP‑LC3 assay and immunofluorescence staining, respectively. The results revealed that treatment with G‑Rg1 enhanced the viability and suppressed the apoptosis of ethanol‑stimulated H9c2 cells. G‑Rg1 also attenuated autophagy and ERS in ethanol‑stimulated H9c2 cells. In addition, the levels of phosphorylated (p)‑protein kinase R (PKR)‑like ER kinase (PERK), p‑eukaryotic translation initiation factor 2a, activating transcription factor 4 (ATF4), CHOP, caspase‑12 and p‑AMPK were downregulated, while the p‑mTOR level was upregulated in ethanol‑stimulated H9c2 cells treated with G‑Rg1.
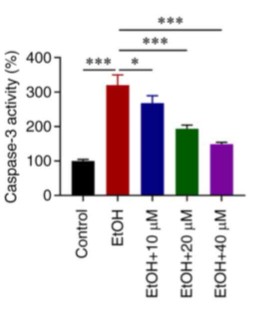
Fig3. Caspase-3 activity in ethanol-stimulated H9c2 cells. (Ge Tian, 2023)
A variety of detection kits are used in proteomics research, which are mainly used for protein extraction, quantification, separation, enrichment, detection and analysis. It is widely used in life science research, medical research, drug development and diagnosis. Creative BioMart offers related kit products for individual processes in this field. If you have any other customized needs, please feel free to contact us!
















