Hippo Signal Pathway
🧪 RHOA-1145H
Source: E.coli
Species: Human
Tag: Non
Conjugation:
Protein Length:
🧪 ITCH-3094H
Source: E. coli
Species: Human
Tag: Non
Conjugation:
Protein Length: 526-903 aa


🧪 CD44-3961H
Source: HEK293
Species: Human
Tag: His
Conjugation:
Protein Length: Gln 21-Pro 220
🧪 RHOA-3975H
Source: Insect Cells
Species: Human
Tag: His
Conjugation:
Protein Length: 1-193 aa

🧪 CD44-10956H
Source: E.coli
Species: Human
Tag: His
Conjugation:
Protein Length: 28-380 aa
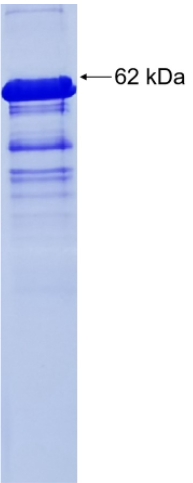
🧪 FRMD5-13005H
Source: E.coli
Species: Human
Tag: GST
Conjugation:
Protein Length: C-term-250a.a.
🧪 CD44-421H
Source: HEK293
Species: Human
Tag: Fc
Conjugation:
Protein Length: Gln 21 - Pro 220
🧪 PPP2R2C-1917H
Source: E.coli
Species: Human
Tag: GST
Conjugation:
Protein Length: C-term-300aa
🧪 RASSF3-2193H
Source: E.coli
Species: Human
Tag: His
Conjugation:
Protein Length: 84-197aa
🧪 RASSF4-2194H
Source: E. coli
Species: Human
Tag: GST
Conjugation:
Protein Length: 1-321 aa

🧪 RASSF7-2195H
Source: E.coli
Species: Human
Tag: His
Conjugation:
Protein Length: 91-292aa
🧪 RASSF8-2196H
Source: E.coli
Species: Human
Tag: GST
Conjugation:
Protein Length: N-term-291aa
🧪 RHOA-2290H
Source: E.coli
Species: Human
Tag: GST
Conjugation:
Protein Length: 1-193aa
🧪 RHOB-2291H
Source: E. coli
Species: Human
Tag: GST
Conjugation:
Protein Length: 1-196 aa

🧪 RHOD-2293H
Source: E.coli
Species: Human
Tag: His
Conjugation:
Protein Length: 1-210aa
Background of Hippo Signal Pathway
What is Hippo Signal Pathway?
Hippo signaling pathway is a highly conserved signaling pathway in the process of evolution, which plays a key role in regulating cell proliferation and apoptosis in multicellular organisms, and has an important influence on maintaining tissue homeostasis and organ size. This pathway was first discovered in fruit flies, and at its core is a kinase cascade involving multiple protein interactions and phosphorylation processes.
Hippo activation is coupled to diverse stimuli, including cell contact, cell stress, and growth factor receptor signaling. These stimuli engage the Hippo pathway via multiple different signaling mechanisms that are cell type and cell context dependent. Notably, distinct mechanisms that control Hippo signaling have been uncovered in hematopoietic, epithelial, and mesenchymal lineages, and in cells exposed to strong inducers of apoptosis.
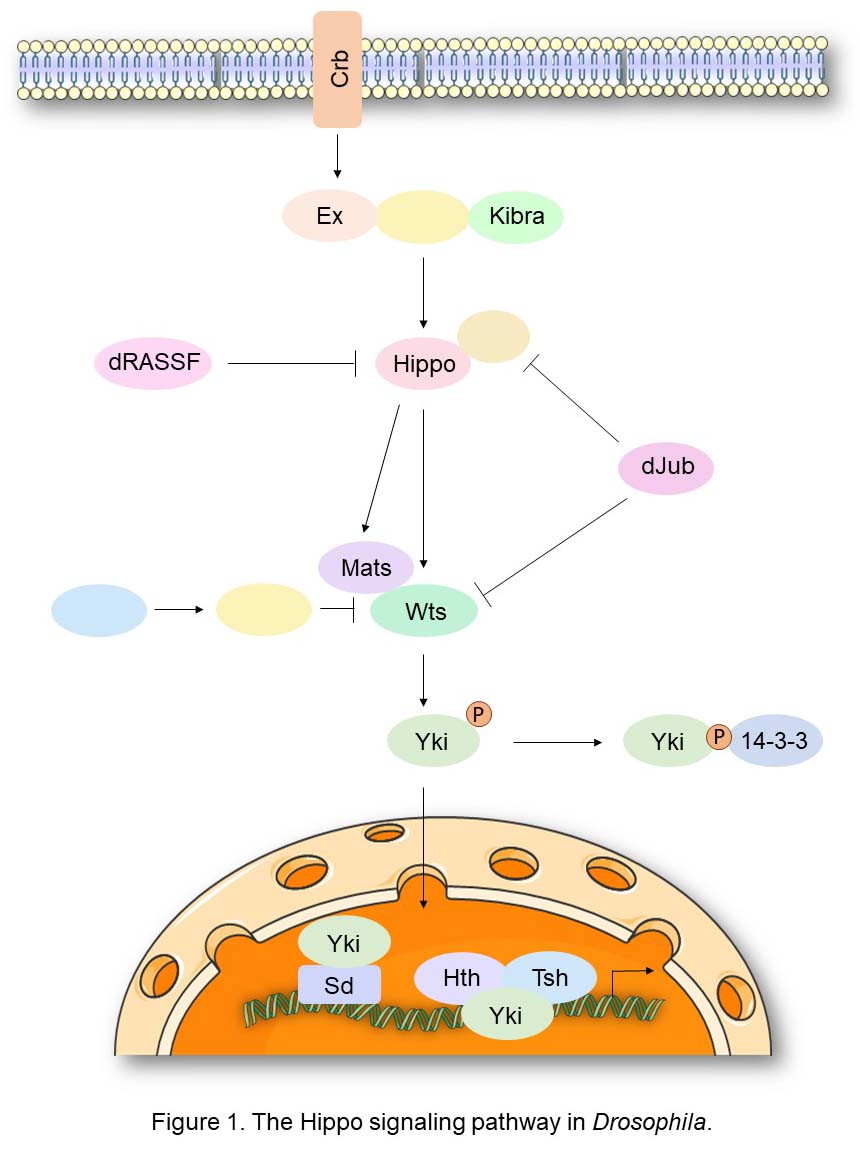
Hippo Signaling in Drosophila
The Hippo signaling pathway was first identified in Drosophila through genetic screens for genes controlling of tissue growth. At the core of this signaling cascade is the Hippo kinase, which functions as a tumor suppressor and regulator of organ size. Hippo binds and phosphorylates the non-catalytic scaffold protein Salvador (Sav), facilitating Hippo-mediated phosphorylation and activation of the Warts (Wts) kinase. The latter acts with the adaptor protein Mats by direct binding. This cascade culminates in the Wts-mediated phosphorylation of the oncogenic transcriptional coactivator Yorkie (Yki). As a result, Yki is bound by 14-3-3 proteins leading its cytoplasmic retention and consequent inactivation. Yki is the critical downstream target of Hippo signaling as the overgrowth phenotypes observed in the Hippo, Sav, or Wts loss-of-function mutants are associated with the constitutive nuclear localization of Yki, and correspondingly Yki inactivation rescues these phenotypes. Yki’s function requires association with TEA-domain (TEAD) family DNA-binding transcription factors, including Scalloped (Sd), Homothorax (Hth), and Teashirt (Tsh). Key target genes activated by Yki include Cyclin E, DIAP1, and the microRNA bantam, which confer increased proliferation and defects in developmental apoptosis.
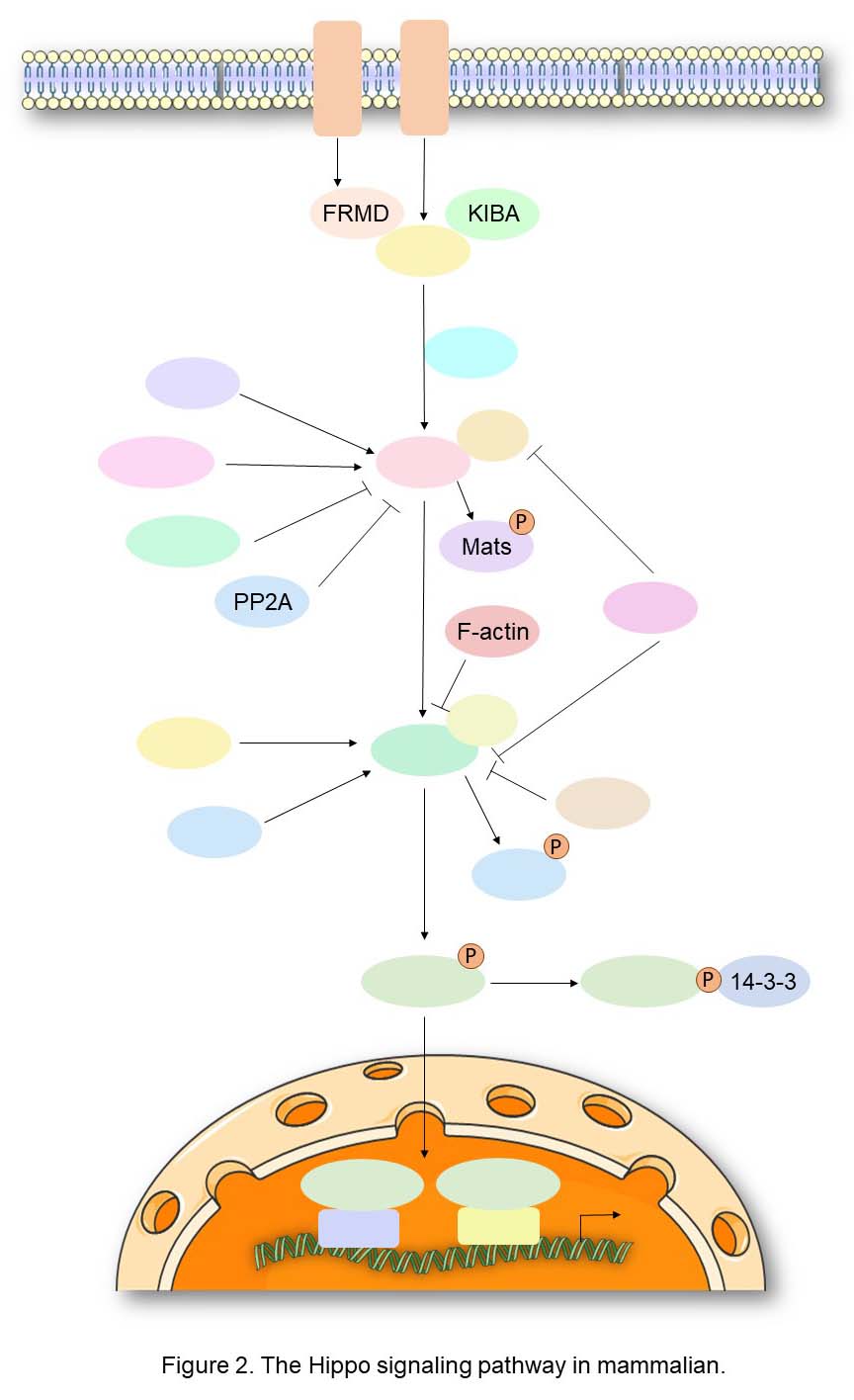
Hippo Signaling in Mammals
All of the core components of the Hippo signaling pathway are conserved and duplicated in mammalian genomes, except for Sav and Yki that are present as single orthologues in vertebrates, (SAV1/WW45 and YAP, respectively), although YAP has a paralogue, TAZ, which is also negatively regulated by this pathway. These components in mammals form a kinase cascade similar to that observed in flies, and several loss-of-function mutant phenotypes observed in Drosophila can be rescued by the expression of their human counterparts. Indeed, the sterile 20-like kinases MST1/STK4 and MST2/STK3, orthologues of the Drosophila Hippo kinase, interact with their regulatory subunit SAV1 to form an active complex that can phosphorylate and activate the kinases LATS1 and LATS2, the direct orthologues of Warts. LATS1, 2 are regulated by the Mats orthologues, MOBKL1A/B (collectively referred to as MOB1), which are also phosphorylated by MST1/MST2 to enhance binding in the LATS1, 2-MOB1 complex. Activated LATS1, 2 phosphorylates the transcriptional coactivator YAP on five different consensus HXRXXS motifs. As for Yki in Drosophila, the phosphorylation of YAP at Ser127 (Ser89 for TAZ) promotes their cytoplasmic retention through 14-3-3 binding. Moreover, phosphorylation of Ser381 primes YAP for subsequent phosphorylation by CK1 in a phosphodegron, thereby promoting the recruitment of the SCFβ-TRCP E3 ubiquitin ligase which catalyzes YAP ubiquitination, ultimately leading to proteosomal degradation of YAP. Unphosphorylated YAP/TAZ primarily localize in the nucleus, where they interact with DNA-binding transcription factors such as TEAD1-4 or SMAD proteins, in order to regulate a transcriptional program that promotes cell survival and proliferation.
Core Components of Hippo Signal Pathway
Merlin and Angiomotin
Merlin contains an N-terminal FERM domain that comprises three subdomains organized into a cloverleaf-like structure, followed by a coiled-coil domain and a charged C-terminal tail. The NF2 allele is alternatively spliced resulting in two predominate forms of the Merlin protein (isoform 1 and 2) that differ at the extreme C-terminus. Merlin and another FERM domain protein, Expanded, were first identified as upstream regulators of Hippo signaling through genetic screens in Drosophila. Merlin regulates the expression and localization of YAP in mammalian cells in a manner similar to what has been observed in flies. In Drosophila, Merlin was shown to form an apical complex with Kibra and together they activate Sav and Wts, leading to Yki phosphorylation and inactivation. Yeast two hybrid and biochemical studies indicated that Merlin directly binds to Sav through its N-terminal FERM domain and Kibra through its C-terminal half.
As members of the Motin protein family, Angiomotin (Amot), Angiomotin-like 1 (AmotL1), and Angiomotin-like 2 (Amotl2) are characterized by a conserved N-terminal glutamine-rich domain, followed by a coiled-coil domain and a C-terminal PDZ-binding motif. Recent work has implicated the Angiomotin family members in the regulation of Hippo-YAP signaling. At least under conditions used for immunoprecipitation, the Angiomotins are arguably one of the strongest binding partners for YAP, as evidenced by four independent studies identifying them as major YAP-associated proteins. The interactions between the Angiomotins and YAP are mediated through PPXY motifs within the N-terminal regions of the Angiomotins and the conserved WW domains of YAP.
MST1/2 Regulation and Substrates
MST1/2 proteins have long been the subject of intense study because of their involvement in stress-induced apoptosis. In this context, MST1/2 are activated by autophosphorylation and caspase-dependent cleavage, which liberates the ~35 KDa N-terminal kinase domain from a C-terminal autoinhibitory domain; active kinase then translocates the nucleus and promotes apoptosis by phosphorylating relevant substrates such as Histone H2B. However, the caspase cleavage site of MST1/2 is not conserved in Hpo, and mutant alleles of hpo that encode an analogous ~35 KDa product lead to loss- rather than gain-of-function mutations in Drosophila.
MST1/2 activity is also regulated by heterodimerization with RASSF family proteins. The human genome encodes six distinct RASSF family members (RASSF1 to -6), each of which contains a Ras association (RA) domain and a C-terminal SARAH domain. Most RASSF family members have been implicated as tumor suppressors, and their expression is frequently inactivated through epigenetic silencing in human cancers. Their tumor suppressor activity can often be attributed, at least in part, to SARAH domain-mediated heter-odimerization with MST1/2. This interaction may activate MST1/2 by targeting MST1/2 to their endogenous activators and substrates or by liberating MST1/2 from their inhibitors (such as the kinase Raf-1).
Accompanying the complex mechanisms of MST1/2 regulation is the complexity of known MST1/2 substrates. Besides the core Hippo pathway components WW45, MOB1 and LATS1/2, MST1/2 have been shown to phosphorylate a diverse array of substrates, such as Histone H2B, transcription factors FOXO1 and FOXO3, and the LATS1/2-related kinases Ndr1/Ndr2, mostly in response to apoptotic stimuli. It will be important to define the contribution of these pathways to the overall tumor suppressor function of MST1/2, especially in the context of intact mammalian tissues.
YAP and TAZ
The Yes-associated protein (YAP) and WW domain-containing transcription regulator 1 (WWTR1, also known as TAZ) are two transcription co-activators that act downstream of the Hippo tumor suppressor pathway. YAP/TAZ regulate expression of a large number of genes that are important in controlling organ size, tumorigenesis, and stem cell functions. The activity of YAP/TAZ is mainly inhibited by LATS kinases of the Hippo pathway. Upon phosphorylation by LATS kinases, YAP/TAZ are sequestered in the cytoplasm and undergo ubiquitination mediated degradation. YAP/TAZ are also inhibited by interaction with cell junction proteins including angiomotin and a -catenin. Moreover, as transcription co-activators, YAP/TAZ need to associate with DNA-binding proteins such as TEAD family transcription factors to induce gene expression. Hence, the activity and specificity of YAP/TAZ in gene expression is also dependent on their nuclear partners.
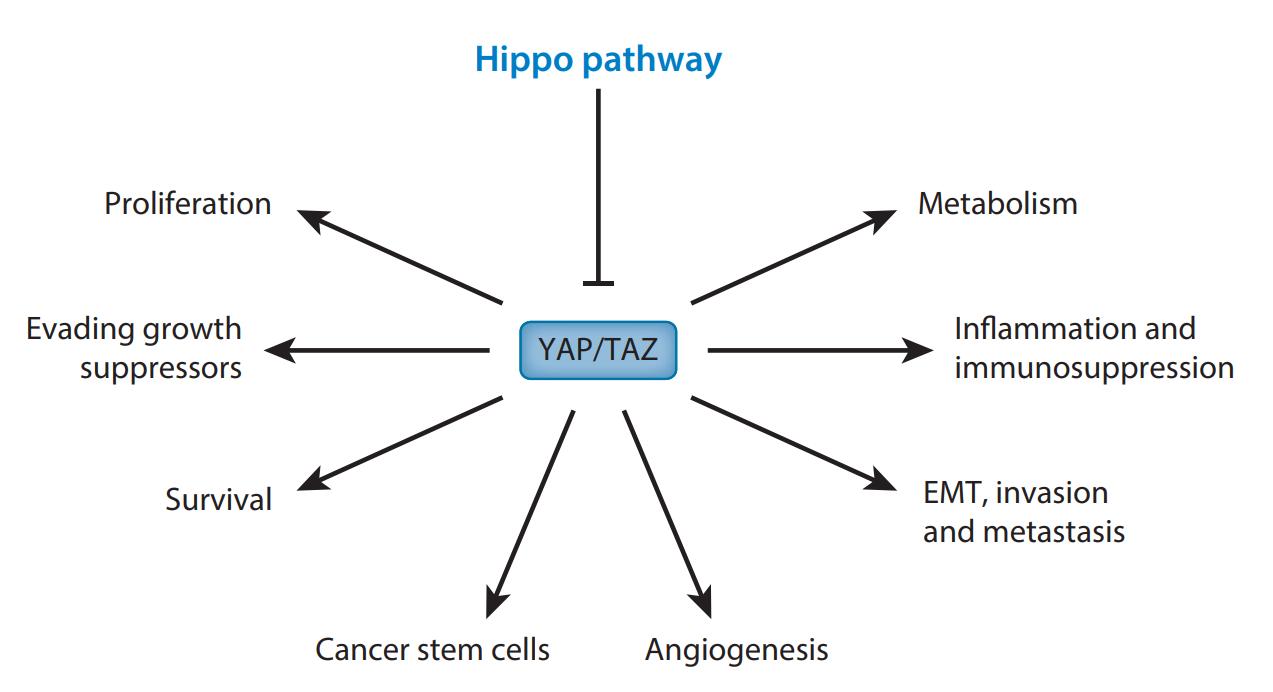
Figure 3. YAP/TAZ and cancer hallmarks. (Shenghong Ma, 2019)
Hippo Signal Pathway Related Diseases
The Hippo signaling pathway plays a key role in regulating biological processes such as cell proliferation, differentiation, organ growth, embryonic development, and tissue regeneration and wound healing. When the Hippo signaling pathway is dysfunctional, it can lead to a variety of diseases, including cancer, eye disease, heart disease, lung disease, kidney disease, liver disease, and immune dysfunction. In addition, the Hippo signaling pathway has been implicated in autoimmune diseases such as rheumatoid arthritis and inflammatory bowel disease. Therefore, targeted therapeutic strategies targeting regulatory components of the Hippo signaling pathway may be promising approaches for treating a broad spectrum of diseases.
Case Study
Case Study 1: Recombinant Human CD44 protein (CD44-3961H)
Due to the heterogeneity and high frequency of genome mutations in cancer cells, targeting vital protumour factors found in stromal cells in the tumour microenvironment may represent an ideal strategy in cancer therapy. However, the regulation and mechanisms of potential targetable therapeutic candidates need to be investigated. Clinically, the results indicate that stromal PTX3 expression correlates with adverse prognostic features and is associated with worse survival outcomes in triple-negative breast cancer (TNBC). The researchers also found that transforming growth factor beta 1 (TGF-β1) induces PTX3 expression by activating the transcription factor CCAAT/enhancer binding protein delta (CEBPD) in stromal fibroblasts. Following PTX3 stimulation, CD44, a PTX3 receptor, activates the downstream ERK1/2, AKT and NF-κB pathways to specifically contribute to the metastasis/invasion and stemness of TNBC MDA-MB-231 cells.

Fig1. The CD44 binding activity of P2rdAD9 or P12rdGR9 was assessed by incubation with CD44, and their interaction was analysed by ELISA. (Yu-Wei Hsiao, 2022)
Case Study 2: Recombinant Human MST1 protein
In flies and mammals, the Hippo pathway controls organ growth, tissue homeostasis, and regeneration. The Hippo pathway can be affected by numerous external and internal factors, which influence the Hippo core kinase cassette. The best understood Hippo core kinase cassette comprises the MST1/2 (aka STK4 and STK3) and LATS1/2 protein kinases. Upon activation the MST1/2 kinases can phosphorylate the LATS1/2 protein kinases and the MOB1 signal transducer protein, thereby supporting the formation of an active MOB1/LATS complex, which appears to be essential for development and tissue growth control. The complexity of MST1 substrates makes it crucial to accurately define and validate direct MST1/2 substrates using in vitro kinase assays. Hence, this study describes the in vitro phosphorylation of recombinant full-length MOB1 by full-length recombinant MST1/2 kinases.
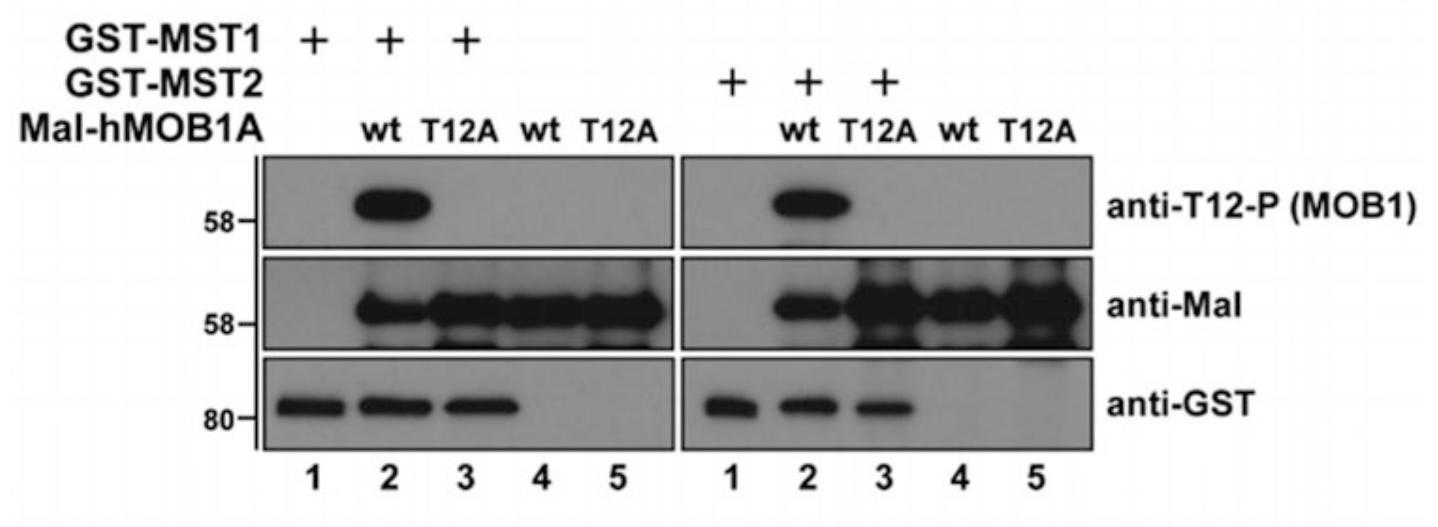
Fig2. Kinase assays using recombinant GST-MST1/2 and Mal-MOB1. (Marta Gomez, 2019)
Case Study 3: Recombinant Human Merlin
Merlin is encoded by the neurofibromatosis type 2 (NF2) gene and is a member of the Band 4.1 protein family. This protein acts as a linker that connects cell surface proteins to the actin cytoskeleton. Defects caused by mutations of the NF2 gene give rise to NF2 disease. However, the role of merlin in human melanoma growth and the mechanism underlying its effect are currently unknown. This study shows that merlin knockdown enhances melanoma cell proliferation, migration, and invasion in vitro and that decreased merlin expression promotes subcutaneous melanoma growth in immunocompromised mice. Concordantly, the researchers find that increased expression of merlin in a metastatic melanoma cell line reduced their in vitro migration and proliferation, and diminished their ability to grow in an anchorage independent manner. Increased merlin expression also inhibits in vivo growth of these melanoma cells. Lastly, they demonstrate that higher merlin levels in human melanoma cells promote the H(2)O(2)-induced activation of MST1/2 Ser/Thr kinases, which are known tumor suppressors in the Hippo signaling pathway.

Fig3. Western blot analysis of endogenous merlin expression in human melanoma cell lines. (Lucas B Murray, 2012)
Related Products
The Hippo pathway was initially discovered in Drosophila melanogaster as a key regulator of tissue growth. It is an evolutionarily conserved signaling cascade regulating numerous biological processes, including cell growth and fate decision, organ size control, and regeneration. Creative BioMart can provide a list of core protein products of the Hippo pathway in mammals consists of a kinase cascade, MST1/2 and LATS1/2, as well as downstream effectors, transcriptional coactivators YAP and TAZ. Please feel free to contact us if you’re interested.