Lipid Transfer Assay
Background
Lipid transfer proteins play a vital role in lipid homeostasis and lipoprotein metabolism. Two key players in this process are Cholesteryl Ester Transfer Protein (CETP) and Phospholipid Transfer Protein (PLTP). These proteins mediate the dynamic exchange of lipids among various lipoproteins, influencing cardiovascular health, metabolic pathways, and the efficacy of lipid-modifying therapies.
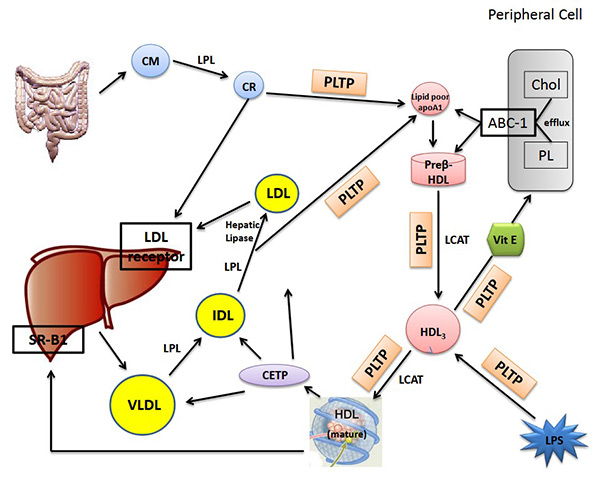
Figure 1. Lipoprotein metabolism and lipid transfer proteins with effects on dietary lipoproteins (chylomicrons), hepatic derived lipoproteins (VLDL, IDL, LDL) and HDL lipoproteins. (Martins and Creegan, 2014).
CETP facilitates the bidirectional transfer of cholesteryl esters and triglycerides between HDL, LDL, and VLDL particles, influencing cholesterol distribution. PLTP is central to phospholipid exchange and HDL remodeling, with implications in metabolic syndromes and insulin resistance. Because of their crucial physiological functions and relevance to drug targets (e.g., statins), assaying CETP and PLTP activity is of significant pharmacological interest.
Creative BioMart offers a specialized Lipid Transfer Assay platform for evaluating CETP and PLTP activity, as well as screening for potential inhibitors. Utilizing a self-quenched fluorescent neutral lipid probe, we provide real-time, quantitative measurement of lipid transfer with exceptional sensitivity and reproducibility. Our platform enables both functional enzymatic analysis and inhibitor drug screening, making it a powerful tool for cardiovascular and metabolic disease research.
What We Offer?
Service Procedure
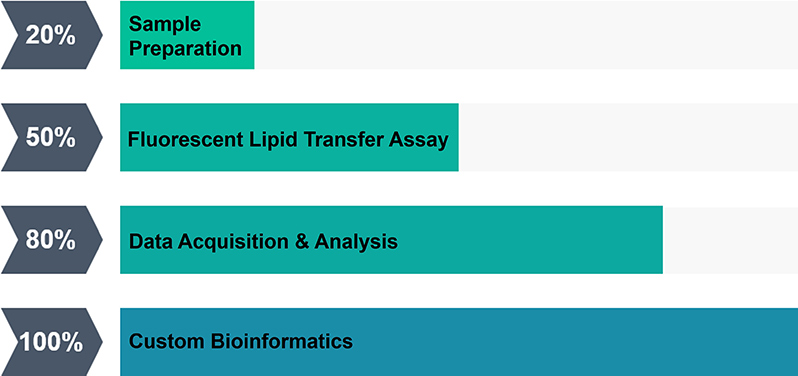

Service Offering
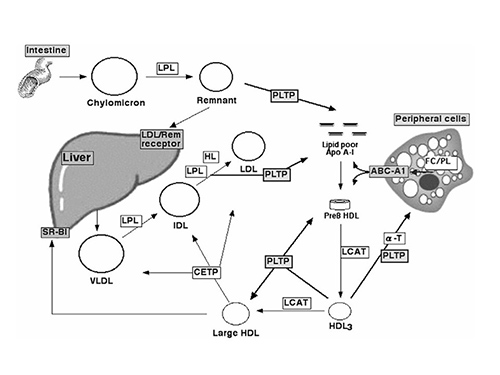 |
CETP & PLTP Activity Assay Our assay uses self-quenched fluorescent neutral lipids to directly measure lipid transfer activity in real time. CETP-mediated cholesteryl ester/triglyceride exchange and PLTP-facilitated phospholipid transfer are quantified by tracking fluorescence recovery, providing accurate kinetic data. Assays are compatible with serum, plasma, and purified proteins. |
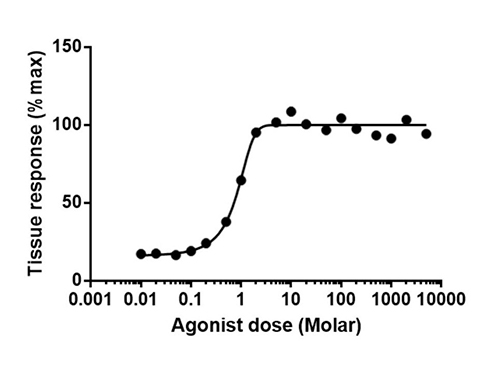 |
Inhibitor Compounds Screening Using the same lipid transfer platform, we evaluate the inhibitory potential of drug candidates against CETP or PLTP. Dose-response curves, IC50 values, and mechanistic insights are generated efficiently with high sensitivity and reproducibility—perfect for hit validation and early-phase lead optimization. |
Service Details
- Assay Format: Fluorescence-based using self-quenched neutral lipid probe
- Readout: Fluorometric intensity proportional to lipid transfer
- Applications: Enzyme activity measurement, inhibitor screening, drug mechanism studies
- Sample Types: Serum, plasma, reconstituted lipoproteins, purified proteins
- Customization: Available for target-specific or high-throughput screening formats
- Turnaround Time: Typically, 5–10 business days
Why Choose Us?
- Advanced Fluorescent Technology: Real-time, quantitative lipid transfer measurement using self-quenched fluorophores—no radiolabels, antibodies, or secondary detection steps required.
- Dual Target Capability: Simultaneously assess CETP and PLTP activity on a single, unified platform with standardized conditions and comparable results.
- Scientific Expertise: Led by experts in lipid metabolism and enzymology with proven experience in assay design, validation, and drug screening.
- Fast & Flexible: Rapid turnaround with customizable assay formats tailored to both basic research and pharmaceutical development needs.
- High Sensitivity & Reproducibility: Optimized fluorescence detection ensures robust, consistent results across experiments with excellent signal-to-noise ratio and lot-to-lot stability.
- High-Throughput Capability: Compatible with 96- and 384-well plate formats for efficient large-scale screening and compound library profiling.
Case Study
* NOTE: We prioritize confidentiality to safeguard our clients’ technology and intellectual property. As an alternative, we present selected published research articles as representative case studies. For details on the assay services and products used in these studies, please refer to the relevant sections of the cited literature.
Case 1: Cholesteryl ester transfer protein (CETP) as a drug target for cardiovascular disease
Schmidt et al., 2021. doi:10.1038/s41467-021-25703-3
This study evaluates the potential of CETP (cholesteryl ester transfer protein) as a therapeutic target for coronary heart disease (CHD) using clinical trial data and Mendelian randomization. Although past CETP inhibitors have failed in trials, the analysis suggests these failures were likely due to compound-specific issues rather than the CETP target itself. Genetic evidence supports that CETP inhibition can lower the risk of CHD, heart failure, diabetes, and chronic kidney disease, but may raise the risk of age-related macular degeneration. Compared to PCSK9 inhibition, which has broader cardiovascular benefits and different side effects, combined targeting of both proteins may yield enhanced therapeutic outcomes.
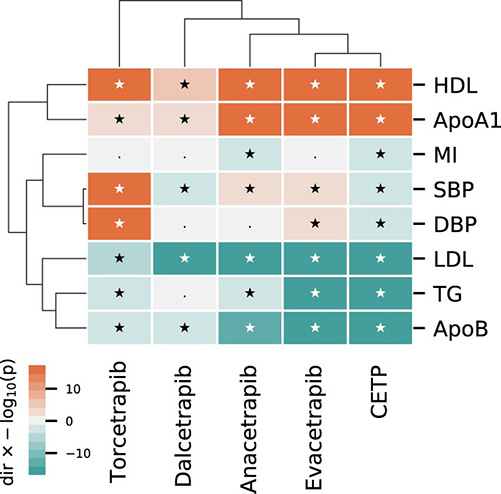
Figure 2. A cluster analysis comparing the on-target Mendelian randomization effect of lower CETP concentration to effects from CETP inhibiting compounds. (Schmidt et al., 2021)
Case 2: DNMT3B-mediated PLTP hypermethylation aggravates vascular dysfunction in diabetic retinopathy via AKT/GSK3β Pathway
Cai et al., 2025. doi:10.1186/s13148-025-01874-4
This study investigates how phospholipid transfer protein (PLTP) affects vascular dysfunction in diabetic retinopathy (DR) and explores its regulation via DNA methylation. Using high-glucose-treated human retinal microvascular endothelial cells and diabetic mice models, researchers found that PLTP expression is reduced due to promoter hypermethylation mediated primarily by DNMT3B. PLTP overexpression reversed high glucose–induced impairments in endothelial cell migration and tube formation. Mechanistically, PLTP promotes phosphorylation of AKT and GSK3β, indicating it supports angiogenesis through the AKT/GSK3β signaling pathway. Overall, abnormal methylation-driven PLTP downregulation contributes to vascular dysfunction in DR.
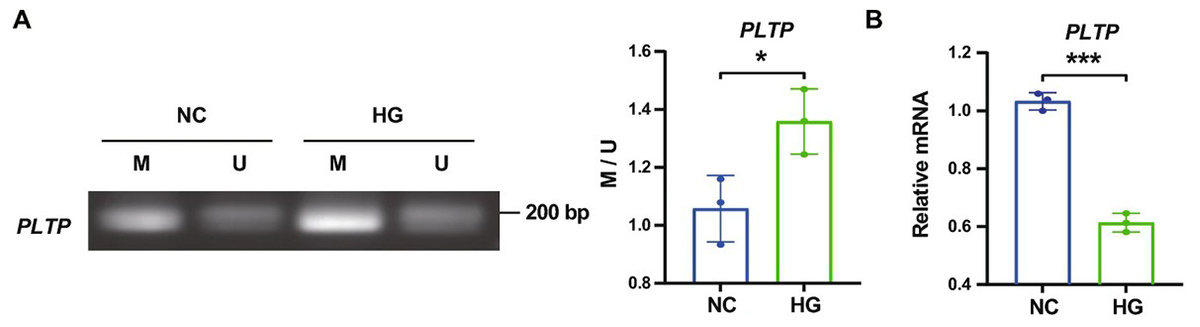
Figure 3. Abnormal PLTP is Involved in Vascular Dysfunction in DR. A. Agarose gel electrophoresis results of MS-PCR products of PLTP in HRMECs (human retinal microvascular endothelial cells) and quantitative analysis of promoter methylation levels. B. PLTP expression in HRMECs. (Cai et al., 2025)
Customer Testimonials
-
"The lipid transfer assay from Creative BioMart provided clean, reproducible CETP activity data, outperforming our in-house system in both speed and cost-efficiency. The platform integrated well with our lipid metabolism screening workflow."
— Senior Scientist, Cardiovascular Drug Discovery Program | Global Pharmaceutical Company
-
"We received thoughtful technical consultation and rapid delivery. The PLTP activity assay was fully compatible with our human plasma samples, enabling smooth data collection in our biomarker validation study."
— Project Manager, Translational R&D | European Biotech Organization
-
"Creative BioMart’s CETP assay consistently generated high-quality, reproducible data that supported key milestones in our small-molecule inhibitor program. Communication with the team was prompt, and timelines were met ahead of expectations."
— Director, Preclinical Pharmacology | Biopharmaceutical R&D Division
-
"The PLTP assay proved instrumental in identifying novel HDL-associated metabolic interactions in our lipidomics research. It’s a solid platform for mechanistic studies in lipoprotein biology."
— Principal Investigator | Academic Medical Center
FAQs
-
Q: What types of samples are compatible with the assay?
A: Our assay supports serum, plasma, purified lipoproteins, or recombinant protein systems. -
Q: Can the assay be adapted for high-throughput screening (HTS)?
A: Absolutely. The platform is scalable for 96- and 384-well formats for screening large compound libraries. -
Q: What fluorophores are used?
A: We use proprietary self-quenched fluorophores optimized for neutral lipid tracking with high signal-to-noise ratios. -
Q: Is inhibitor profiling included?
A: Yes. We offer inhibitor screening services with IC₅₀ determination and optional mechanistic follow-up studies. -
Q: How is data delivered?
A: Reports include raw fluorescence data, normalized activity values, and dose-response curves (if applicable).
Resources
Related Services
- Electron Transfer Quenching Platform
- Protein Microarray Service
- Protein Pathway Profiling
- Bioanalytical Assay Services
Related Products
References:
- Cai C, Gu C, Meng C, et al. DNA hypermethylation of PLTP mediated by DNMT3B aggravates vascular dysfunction in diabetic retinopathy via the AKT/GSK3β signaling pathway. Clin Epigenet. 2025;17(1):82. doi:10.1186/s13148-025-01874-4
- Martins IJ, Creegan R. Links between insulin resistance, lipoprotein metabolism and amyloidosis in Alzheimer’s Disease. Health. 2014;06(12):1549-1579. doi:10.4236/health.2014.612190
- Schmidt AF, Hunt NB, Gordillo-Marañón M, et al. Cholesteryl ester transfer protein (CETP) as a drug target for cardiovascular disease. Nat Commun. 2021;12(1):5640. doi:10.1038/s41467-021-25703-3
Contact us or send an email at for project quotations and more detailed information.
Quick Links
-

Papers’ PMID to Obtain Coupon
Submit Now -

Refer Friends & New Lab Start-up Promotions

