What is DAO Protein
D-amino-acid Oxidase, commonly referred to as DAO, is a type of protein that plays a critical role in the human body, particularly by breaking down D-amino acids. This protein family is ubiquitous in many organisms ranging from bacteria to mammals, implicating their fundamental role in cellular activities.
The DAO protein was first isolated and identified in 1935, by two Japanese scientists, Kotaro Suzuki and Rohdoh Udenfriend, while they were studying pig kidney. The gene encoding DAO protein, the DAAO gene, primarily resides in humans on the 12q24 locus of chromosome 12. Two exons in this gene are responsible for the coding and production of the DAO protein.
The structure of DAO protein is fairly complex, encompassing 347 amino acids and approximately 39kDa. It has primary, secondary, tertiary, and quaternary structures like all other proteins. The tertiary structure of DAO comprises a highly conserved flavin adenine dinucleotide (FAD) binding domain, implicating FAD as a vital cofactor for DAO activation.
Function of DAO protein
The primary function of the DAO protein is to oxidize D-amino acids, such as D-serine and D-alanine, to their corresponding α-keto acids. DAO catalyzes the oxidative deamination to produce α-keto acids, ammonia, and hydrogen peroxide. Earlier, the enzymatic activity of DAO was thought to be involved merely in detoxification processes. However, recent studies suggest that mammalian DAO is involved in multiple physiological and developmental processes, playing an essential role in signal transduction.
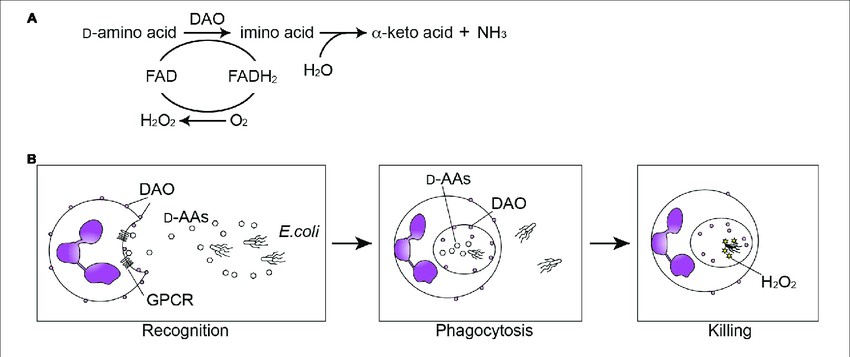
Fig1. Emerging Role of D-Amino Acid Metabolism in the Innate Defense (Sasabe, Jumpei & Suzuki, Masataka. 2018)
DAO protein related signal pathway
The DAO protein plays a pivotal role in various signal pathways. A critical pathway includes the modulation of N-methyl-D-aspartate (NMDA) receptors' activity, particularly involved in the glutamatergic neurotransmission system in the brain. DAO equilibrium exposure to D-serine, an endogenous co-agonist of NMDAR, determines the proper functioning of this receptor in physiological and pathological conditions. Hence, DAO serves as the regulator of NMDA receptor activity, ensuring the proper neuronal functions and maintaining appropriate Glutamatergic transmission.
DAO protein related diseases
While DAO is central to maintaining homeostasis, any anomalies in its regulation could lead to a furnished array of diseases. Studies indicate a connection between aberrant DAO function and entire diseases, including neurodegenerative disorders. DAO dysfunctions contribute to the altered schizophrenia symptoms, leading to cognitive impairment and negative symptoms due to the enhanced NMDA receptor activity. Moreover, DAO over activity links to amyotrophic lateral sclerosis by triggering motor neuron degeneration. DAO activity also implicates in pulmonary diseases and cancer development. These findings reflect the critical role of DAO in pathological conditions, extending the diagnostic and therapeutic horizons.
DAO protein's applications in biomedical
The DAO protein has garnered substantial attention in biomedical applications due to the consequences of its dysfunction. The protein is identified as a clinical target in developing therapeutic interventions for various diseases. Researchers are working on controlling DAO activity to manage diseases related to its dysfunction. For instance, DAO inhibitors can potentially improve schizophrenia symptoms by enhancing D-serine levels, ultimately upsurging NMDA receptor activity. Additionally, DAO can function as a potential biomarker for detecting certain types of cancers early.
In conclusion, the functions and involvements of DAO proteins in various cellular activities and disease conditions are extensive and fundamental. Since its discovery, there has been an increase in understanding its functional and genetic mechanisms, its role in disease development, and its clinical significance. The DAO protein will continue to be an arena of intense investigation, with promising perspectives for developing tailored therapeutic strategies for numerous debilitating diseases.
Our Featured Products
| Cat.No. | Product Name | Species | Source (Host) | Tag |
|---|---|---|---|---|
| DAO-476H | Recombinant Human D-amino-acid Oxidase, His-tagged | Human | E.coli | His |
| DAO-1181H | Recombinant Human DAO Protein, His-SUMO-tagged | Human | E.coli | His/SUMO |
| DAO-11821H | Recombinant Human DAO, GST-tagged | Human | E.coli | GST |
| DAO-4302M | Recombinant Mouse DAO Protein | Mouse | Mammalian Cell | His |
| DAO-2198M | Recombinant Mouse DAO Protein, His (Fc)-Avi-tagged | Mouse | HEK293 | His (Fc)-Avi |
| DAO-194C | Recombinant Cynomolgus Monkey DAO Protein, His (Fc)-Avi-tagged | Cynomolgus Monkey | HEK293 | His (Fc)-Avi |
| CED055 | Active Porcine D-Amino Acid Oxidase | Pig | Porcine Kidney | N/A |
Reference
- Sasabe, Jumpei & Suzuki, Masataka. (2018). Emerging Role of D-Amino Acid Metabolism in the Innate Defense. Frontiers in Microbiology. 9. 10.3389/fmicb.2018.00933.

