Metal/Element Test Service
Background on Heavy Metal and Element Testing in Food Safety Monitoring
In recent years, heavy metal contamination has become a growing global concern due to the intensification of mining, smelting, and industrial manufacturing. These activities can introduce hazardous metals into the soil, water, and air—eventually accumulating in the food chain at levels thousands of times higher than the original concentration.
Once inside the human body, heavy metals can disrupt metabolic processes by binding to critical proteins and enzymes or accumulating in organs, leading to chronic toxicity. Symptoms of heavy metal poisoning may include respiratory distress, gastrointestinal upset, neurological damage, and, in severe cases, birth defects or carcinogenesis.
To mitigate these risks, regulatory agencies worldwide have established strict maximum residue limits (MRLs) and mandate routine testing for heavy metal contaminants in food and health-related products.
At Creative BioMart , we offer a comprehensive portfolio of Metal and Elemental Testing Services to ensure the safety and compliance of food, agricultural products, cosmetics, and environmental samples. Our testing covers both essential minerals (e.g., calcium, iron) and toxic heavy metals (e.g., mercury, lead, cadmium, arsenic, chromium), utilizing advanced analytical instrumentation for precise, reproducible results.
Overview of Our Elemental Analysis Service
Service Procedure

Elements We Test
We support a wide range of metal and elemental testing—covering both essential minerals and toxic heavy metals commonly regulated in food, healthcare, and environmental industries. Whether you're verifying nutritional content or screening for contaminants, our service is designed to meet international safety and compliance standards.
-
Lead

-
Cadmium

-
Mercury

-
Arsenic

-
Chromium

-
Iron

-
Zinc

-
Copper

-
Selenium

-
Nickel

-
Aluminum

-
Tin

-
Cobalt

-
Manganese

-
Magnesium

-
Calcium

-
Sodium

-
Potassium

Sample Types We Support
Creative BioMart accepts a wide range of sample types to meet the diverse testing needs of food safety, healthcare, and environmental industries. Our metal/element testing services are compatible with:
- Food & Beverages : Raw materials, processed food, dairy, edible oils, grains, dried fruits, teas, beverages, etc.
- Dietary Supplements : Powders, tablets, capsules, and herbal extracts.
- Pharmaceuticals : Active ingredients, excipients, and finished dosage forms.
- Cosmetics & Personal Care Products : Creams, serums, lotions, and raw cosmetic materials.
- Environmental Samples : Soil, sludge, sediment, drinking water, wastewater, irrigation water.
- Agricultural Inputs : Fertilizers, animal feed, and pesticides.
- Industrial Materials : Packaging materials, metal alloys, and raw industrial inputs.
Testing Method
At Creative BioMart, we employ advanced analytical platforms including Atomic Absorption Spectrometry (AAS) and Inductively Coupled Plasma Mass Spectrometry (ICP-MS). These techniques enable precise quantification of trace elements, even at parts-per-billion (ppb) levels, ensuring sensitivity, accuracy, and reproducibility for every test.
-

Atomic absorption spectrometer
-

Inductively coupled plasma mass spectrometer
Quality Assurance & Control
At Creative BioMart, we follow a strict quality assurance protocol to ensure high accuracy, precision, and reproducibility in all elemental analyses. Our QA measures include:
- Certified Reference Materials (CRMs): Each batch includes standards traceable to recognized authorities (e.g., NIST, ISO).
- Internal Standards & Spikes: Used to monitor matrix effects and instrument stability in real-time.
- Blank Controls: Ensure no background contamination or carryover affects results.
- Duplicate & Parallel Testing: Selected samples are re-tested to verify consistency.
- Method Validation: All methods are validated for specificity, sensitivity, linearity, accuracy, and recovery.
- Instrument Calibration & Maintenance: Regular calibration using multi-point standards and scheduled preventative maintenance ensure optimal performance.
- Trained Analysts: Our testing is performed by experienced chemists with expertise in trace metal analysis and regulatory compliance.
Advantages of Our Elemental Analysis Service
- State-of-the-Art Equipment : ICP-MS and AAS ensure highly sensitive detection of both trace and bulk elements.
- Regulatory-Ready Reports : Fully compliant with global standards including FDA, EU, GB, Codex, and WHO guidelines.
- Wide Sample Compatibility : We test food, supplements, cosmetics, environmental matrices, and herbal materials.
- Fast Turnaround & High Accuracy : Delivering reliable, reproducible results in as little as 5–7 working days.
- Robust Quality Control : Every test includes internal standards, certified reference materials, and strict validation protocols.
- Expert Technical Support : Our analysts provide consultative support on sample prep, compliance interpretation, and corrective actions if needed.
Case Studies on Heavy Metal Detection
* NOTE: We prioritize confidentiality to safeguard our clients’ technology and intellectual property. As an alternative, we present selected published research articles as representative case studies. For details on the assay services and products used in these studies, please refer to the relevant sections of the cited literature.
Case 1: Lead content in canned crab assessed by AAS
Agustina et al. , 2021. doi:10.1088/1755-1315/679/1/012012
Canned crab products are at risk of contamination by heavy metals like lead due to aquatic pollution. Using Atomic Absorption Spectrophotometry (AAS) per SNI 2354.5:2011, lead (Pb) levels were tested in canned crabs at BPMHP Semarang. Results showed Pb levels were below the safety threshold of 0.5 mg/kg (SNI 6929:2016), indicating the products are safe for consumption. Minimizing heavy metal residues remains essential for public health and food safety.
Table 1. Average of lead content of heavy metal in canned crab samples. (Agustina et al. , 2021)
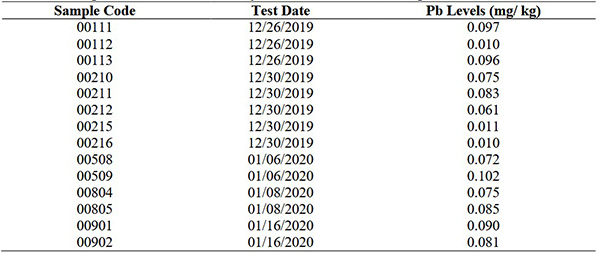
Case 2: Assessment of arsenic, vanadium, mercury, and cadmium in food and drug packaging
Mukhi et al. , 2024. doi:10.12688/f1000research.121473.3
This study investigated heavy metal contamination in 13 types of food and drug packaging materials commonly used in India. Using rapid assays and ICP-OES, researchers detected arsenic, vanadium, mercury, and cadmium—often at levels exceeding European safety limits. Mercury and arsenic were the most frequently found, while vanadium was least common. The findings highlight potential health risks posed by packaging materials and underscore the need for stricter monitoring to ensure consumer safety.
Table 2. Concentrations in parts per million (ppm) of heavy metals detected in study materials. (Mukhi et al ., 2024)
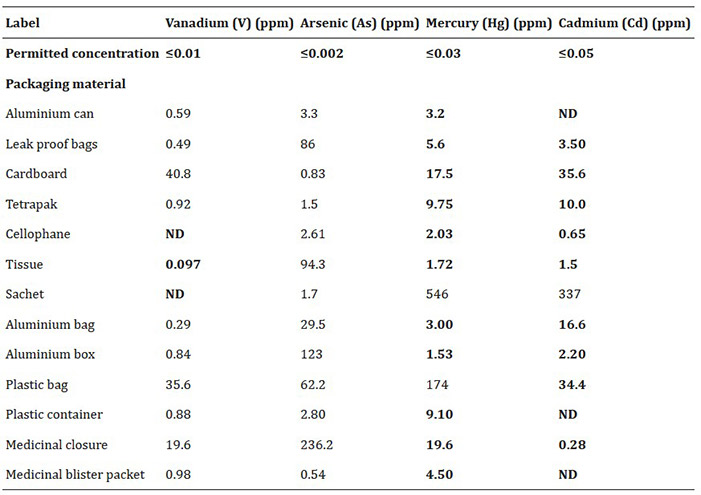
ND = not detected. Bold values indicate values of heavy metals in packaging materials below the or under acceptable amount that is within the given range and not higher than the prescribed limit.
Client Testimonials on Our Metal Element Test Service
“We needed urgent testing for lead and mercury in a batch of imported dried fruits. Creative BioMart provided fast turnaround and clear regulatory comparison data in the report. Their responsive team helped us meet our export deadline without a hitch.”
— Supply Chain Manager | Global Food Distributor
“Creative BioMart ran comprehensive elemental profiling for our dairy-based formula, covering calcium, iron, zinc, and trace metals. Their attention to detail in sample prep and cross-contamination prevention really stood out during our audit review.”
— Formulation Scientist | Infant Nutrition Company
“We required multi-element testing in cosmetic products—including chromium, cobalt, and nickel—to meet REACH and GB compliance. The team at Creative BioMart was not only technically sharp but also highly responsive to regulatory queries.”
— Regulatory Affairs Lead | Personal Care Brand
“Our company faced recurring issues with aluminum residues in a food additive. Creative BioMart’s analysis pinpointed the source with high sensitivity, and their follow-up recommendations helped us tighten our process controls effectively.”
— Head of Process Engineering | Food Ingredient Manufacturer
FAQs on Our Metal/Element Test Service
-
Q: What types of samples can you test for metal/element content?
A: We can test a wide range of sample types, including food, beverages, dietary supplements, agricultural products, cosmetics, and environmental samples like soil or water. -
Q: Which testing technologies do you use, and how accurate are they?
A: We use advanced techniques such as ICP-MS and AAS, offering high sensitivity and precision—even at parts-per-billion (ppb) levels. All results are validated against certified reference materials. -
Q: Can you test both nutritional elements and toxic heavy metals?
A: Yes. Our service covers essential minerals like calcium, iron, and zinc, as well as toxic metals such as lead, mercury, cadmium, arsenic, and chromium. -
Q: How long does the testing process take?
A: Our standard turnaround is 5–7 working days, with expedited services available upon request for urgent regulatory or quality control needs. -
Q: Do your results meet international regulatory standards?
A: Absolutely. Our reports are designed to comply with standards from FDA, EU, Codex, GB (China), and other global food and drug safety authorities. -
Q: What sets you apart from other testing providers?
A: In addition to our advanced instrumentation and experienced analysts, we offer:- Comprehensive test coverage
- Clear regulatory-ready reports
- Strict quality control protocols
- Fast and flexible service options
Resources
Related Services
- Food Additives Test Service
- Fungal Toxin Test Service
- Microbiological Test Service
- Pesticide & Veterinary Residue Test Service
- Nutritive Index Test Service
Related Products
References:
- Agustina M, Mulyono, Tjahjaningsih W. Assessment of heavy metal lead (Pb) contents in canned crab products by atomic absorption spectrophotometry (AAS). IOP Conf Ser: Earth Environ Sci . 2021;679(1):012012. doi:10.1088/1755-1315/679/1/012012
- Mukhi S, Rukmini MS, Ajay Manjrekar P, Iyyaswami R, H. S. A ssessment of arsenic, vanadium, mercury, and cadmium in food and drug packaging. F1000Res . 2024;11:648. doi:10.12688/f1000research.121473.3
Contact us or send an email at for project quotations and more detailed information.
Quick Links
-

Papers’ PMID to Obtain Coupon
Submit Now -

Refer Friends & New Lab Start-up Promotions

