Cell: the “Protein backbone” of the Yeast Telosome
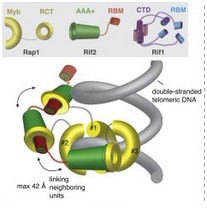
A model of the Rap1-Rif1-Rif2 Assembly at Telomeres
Researchers have made a molecular, biochemical, and functional dissection on the protein backbone at the core of the yeast telosome.
Repeated sequences form the genome as the sequence motif links sequence motif as well as binds proteins. These proteins will control transcription factors in the condition of transcription absence. This seems to be inconsistent. Now researchers have uncovered the situation in a study on telomeres.
We know that telomeres consist of DNA repeats and locate at the ends of chromosomes. With each cell division, the DNA strand of telomere end would shorten a little. There are several processes that can ensure both DNA strands being maintained and the loss of telomeric sequences through unwanted degradation minimizing. While the protective capping of the telomeric ends with a set of dedicated proteins can be used.
The protein cap of the telomere is the telosome. Through structure analysis with x-ray, they found the common transcription factor Rap1, which binds to the telomere repeats, interacts with two proteins called Rif1 and Rif2 to form a higher-order structure that protects the telomeres. In a VELCRO-like manner, up to 20 Rap1 molecules sitting in an array on the DNA repeats are tightly linked to each other through the Rif proteins. Only once this higher-order structure is in place are chromosomal ends protected from degradation thus allowing the cell to sidestep death.
The research result was published on Cell on June 6, 2013.
Article Link: Cell: the “Protein backbone” of the Yeast Telosome

