The Structural Secrets of Actin Nucleation was Found
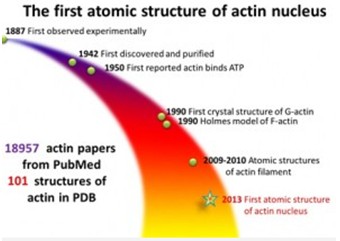
Actin, a sturdy-like filament, helping cells maintain their shape, is an important factor for human. The filaments, named F-actin, were first discovered in late 1900s and been broadly studied since then. F-actin is composed by G-actin, which self-assemble end-to-end. But people are not clearly aware of the assembly process.
F-actin consists of many monomers. During the assembly process, these monomers come together to form the nucleus of a filament in a very efficient way.
In one study targeting to find the structure of the actin nucleus, researchers applied x-ray crystallography. Though the result failed their purpose, they created two mutant versions of G-actin that could nucleate but not polymerize with one could bind normally on top but not on bottom, and the other could bind normally on bottom but not on top, unlike the normal binding which both bind top and bottom.
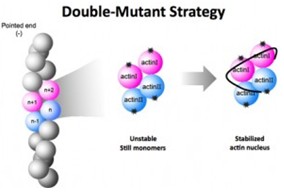
Next, they cultured eligible crystal samples with a two-stage method. The first stage used high levels of super-saturation to spur initial crystal formation. The second stage used a process, seeding to transfer the newly formed crystals to another medium where they could grow large enough for examination. After all settled, they begun to analyze and found the atomic arrangement of each atom in the nucleated, dual-mutant pair.
Thus the process how actin nucleation starts was observed. The finding gathers great benefit for following studies, such as cancer and other diseases.
Article Link: The Structural Secrets of Actin Nucleation was Found

