What is CSF2 Protein
The CSF2 protein, also acknowledged as the granulocyte-macrophage colony-stimulating factor (GM-CSF), is encoded by the CSF2 gene in humans. This crucial cytokine is responsible for stimulating the growth and differentiation of hematopoietic progenitor cells into granulocytes and monocytes, primarily aiding in the making and activation of white blood cells. It plays an indispensable role in immune responses, inflammation, and the functioning of mature granulocytes and macrophages.
CSF2 protein related signal pathway
The multifaceted role of the CSF2 protein also extends to its association with diverse cellular signaling pathways. Signal transduction through the CSF2 protein is predominantly unfolded via the Janus kinase/signal transducer and activator of transcription (JAK/STAT) pathway. The JAK-STAT pathway is a primary signaling mechanism, responsible for various cytokines and growth factors. When CSF2 binds to its receptor on cell surfaces, it triggers the activation of JAK2, which subsequently activates STAT5. Oncodrivers such as activated JAK/STAT can stimulate uncontrolled cell proliferation, making the CSF2/JAK/STAT signaling pathway a potential target for anticancer therapeutic strategies.
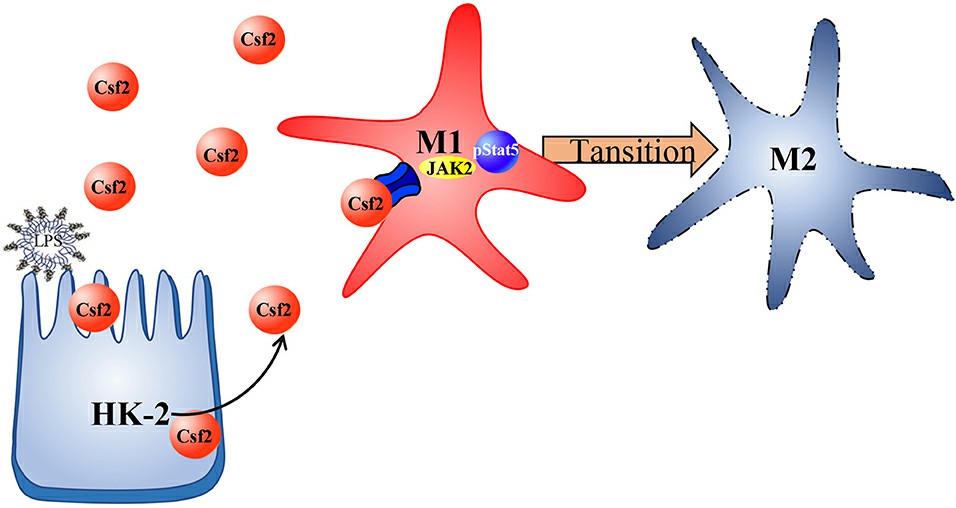
(Li, Y. et al.2020)
CSF2 protein related diseases
Apart from its function in normal biological processes, aberrations in the CSF2 gene have also been implicated in a range of diseases. A salient example is notably Pulmonary Alveolar Proteinosis (PAP), an uncommon condition where mutations in CSF2 lead to the accumulation of lipid protein complexes in the alveoli, impairing gas exchanges in the lungs. The disease, largely driven by CSF2 or GM-CSF antibodies which neutralize the production or functions of the CSF2 protein, highlights the profound clinical significance of CSF2.
CSF2 protein's applications in biomedical
The CSF2 protein also impinges on conditions such as rheumatoid arthritis, where high levels of GM-CSF are present reflecting the severity of the condition. On the other hand, elevated production of the protein has also been perceived in cancer cases. Particularly, certain types of tumors like melanoma and lung cancer exhibit increased levels of GM-CSF, suggesting that it may facilitate tumor progression and resistance to therapies.
Conversely, the CSF2 protein can be harnessed constructively in biomedical applications. Given its capacity to stimulate the production and function of immune cells, GM-CSF has been employed as an adjuvant in vaccine formulations. By augmenting the immunogenicity of vaccines, it potentially enhances a vaccine's effectiveness.
Moreover, recombinant GM-CSF is being used as a therapeutic agent in a variety of clinical settings. It has been approved for use in patients with severe neutropenia, to accelerate bone marrow recovery following chemotherapy or bone marrow transplant. Further, studies pushing the envelope of biomedical research indicate that manipulating the CSF2 protein could hold promising potential for developing targeted disease interventions, especially in cancer therapeutics.
In conclusion, the CSF2 protein or GM-CSF is indispensable to human health and disease. Understanding its intricate roles allows us to appreciate its potential impact on various biomedical applications, particularly in the development of innovative therapeutic strategies. Nevertheless, the precise mechanisms of the CSF2 protein interaction with different diseases remain to be fully elucidated, warranting further research. An in-depth exploration of these mechanisms will help to unlock new avenues for disease management, potentially transforming the future of medical treatments.
Our Featured Products
| Cat.No. | Product Name | Species | Source (Host) | Tag |
|---|---|---|---|---|
| CSF2-486H | Recombinant Human CSF2, Fc tagged | Human | Human Cell | Fc |
| CSF2-210H | Recombinant Human CSF2 Protein, non-tagged, Biotinylated | Human | HEK293 | N/A |
| Csf2-609M | Active Recombinant Mouse Csf2 Protein | Mouse | E.coli | Tag Free |
| Csf2-4346R | Recombinant Rat Csf2 Protein | Rat | Yeast | N/A |
| CSF2-4386F | Recombinant Feline CSF2 Protein | Feline | Yeast | N/A |
| CSF2-4343O | Recombinant Ovine CSF2 Protein | Ovine | Yeast | N/A |
| CSF2-106S | Recombinant Swine CSF2 | Swine | Yeast | N/A |
| CSF2-1729C | Recombinant Chicken CSF2 | Chickein | Mammalian Cell | His |
Reference
- Li, Y., Zhai, P., Zheng, Y., Zhang, J., Kellum, J. A., & Peng, Z. (2020). Csf2 Attenuated Sepsis-Induced Acute Kidney Injury by Promoting Alternative Macrophage Transition. Frontiers in Immunology, 11, 509465. https://doi.org/10.3389/fimmu.2020.01415

