Glutamate Receptors
Related Symbol Search List
Immunology Background
About Glutamate Receptors
Glutamate receptors are a type of receptor protein found in the central nervous system, specifically in the brain, that respond to the neurotransmitter glutamate. Glutamate is the most abundant excitatory neurotransmitter in the brain and plays a crucial role in various physiological processes, including learning, memory, and synaptic plasticity.
There are two primary types of glutamate receptors: ionotropic receptors and metabotropic receptors. Ionotropic receptors directly control the flow of ions across the cell membrane, whereas metabotropic receptors work through intracellular signaling pathways.
Ionotropic glutamate receptors, also known as ligand-gated ion channels, are composed of several subunits that form a pore in the cell membrane. When glutamate binds to the receptor, the channel opens, allowing ions such as calcium, sodium, and potassium to flow into or out of the cell. This ion influx leads to changes in the neuron's electrical potential and can trigger various cellular processes, including the generation of action potentials.
There are three major subtypes of ionotropic glutamate receptors: NMDA (N-methyl-D-aspartate) receptors, AMPA (α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid) receptors, and kainate receptors. NMDA receptors are involved in synaptic plasticity and are critical for learning and memory processes. AMPA receptors mediate fast synaptic transmission and are essential for basic neuronal functions. Kainate receptors have diverse functions and are present in different brain regions.
Metabotropic glutamate receptors, on the other hand, do not directly control ion flow but instead activate intracellular signaling cascades through G proteins. There are three classes of metabotropic receptors: Group I, Group II, and Group III. These receptors modulate neurotransmission, synaptic plasticity, and neuronal excitability by activating various intracellular pathways.
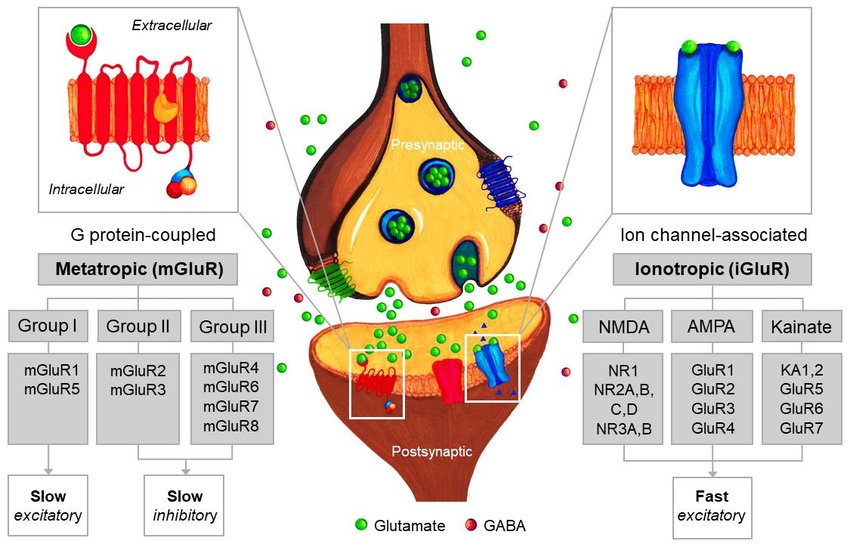 Fig.1 4 Glutamate receptor types. GluR: glutamate receptor; NR: NMDA receptor subtype; KA: kainate receptor subtype. (Tarland A E, 2018)
Fig.1 4 Glutamate receptor types. GluR: glutamate receptor; NR: NMDA receptor subtype; KA: kainate receptor subtype. (Tarland A E, 2018)
Biological Functions of Glutamate Receptors
Excitatory Neurotransmission: Glutamate is the main excitatory neurotransmitter in the brain, and glutamate receptors are responsible for mediating excitatory synaptic transmission. Glutamate receptors allow the influx of calcium and sodium ions, leading to depolarization of the postsynaptic neuron and signal amplification.
Synaptic Plasticity: Glutamate receptors are closely associated with synaptic plasticity, which is the ability of synapses to strengthen or weaken their connections. The two main types of glutamate receptors involved in synaptic plasticity are the N-methyl-D-aspartate (NMDA) receptor and the alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptor. Activation of NMDA receptors is critical for long-term potentiation, which is a process that underlies learning and memory.
Learning and Memory: Glutamate receptors, particularly NMDA receptors, are essential for learning and memory processes. The activation of NMDA receptors leads to the strengthening of synaptic connections and the formation of new synapses, which are thought to be the cellular basis of learning and memory.
Neuronal Development: Glutamate receptors are involved in various aspects of neuronal development. They regulate processes such as cell migration, axon guidance, dendritic arborization, and synaptogenesis. Glutamate receptors also play a role in neurogenesis, which is the generation of new neurons in the adult brain.
Motor Control: Glutamate receptors in the motor cortex and basal ganglia are crucial for motor control. They regulate the excitability of motor neurons and coordinate motor movements. Dysfunction of glutamate receptors in these areas can lead to motor disorders such as Parkinson's disease and Huntington's disease.
Application Fields of Glutamate Receptors
Neurotransmission and Synaptic Plasticity: Glutamate receptors, particularly the ionotropic glutamate receptors, such as NMDA (N-methyl-D-aspartate), AMPA (α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid), and kainate receptors, mediate fast excitatory neurotransmission in the CNS. They are crucial for synaptic plasticity, including long-term potentiation (LTP) and long-term depression (LTD), which underlie learning and memory processes.
Neurodegenerative Diseases: Glutamate excitotoxicity, which refers to excessive glutamate signaling leading to neuronal damage or death, is implicated in various neurodegenerative diseases, including Alzheimer's disease, Parkinson's disease, and Huntington's disease. Understanding the role of glutamate receptors in these conditions can provide insights into disease mechanisms and potential therapeutic targets.
Stroke and Ischemic Injury: During stroke or ischemic events, the excessive release of glutamate can lead to excitotoxicity and neuronal damage. Glutamate receptors, particularly NMDA receptors, play a crucial role in the pathophysiology of ischemic brain injury. Research on glutamate receptors in this context can contribute to the development of neuroprotective strategies and therapeutic interventions.
Epilepsy: Abnormal glutamate signaling and alterations in glutamate receptor function have been implicated in epilepsy, a neurological disorder characterized by recurrent seizures. NMDA and AMPA receptors have been particularly studied in relation to epilepsy. Investigating glutamate receptors can provide insights into the underlying mechanisms of epileptic seizures and aid in the development of antiepileptic drugs.
Pain Processing: Glutamate receptors, especially NMDA receptors, are involved in the transmission and modulation of pain signals in the CNS. Glutamate-mediated synaptic transmission in pain pathways contributes to the development and maintenance of chronic pain conditions. Targeting glutamate receptors has been explored as a potential approach for pain management.
Psychiatric Disorders: Glutamate receptors have been implicated in various psychiatric disorders, including schizophrenia, depression, and addiction. Dysregulation of glutamate signaling, particularly NMDA receptor dysfunction, has been proposed as a contributing factor in the pathophysiology of these disorders. Studying glutamate receptors can provide insights into the mechanisms underlying these conditions and potential targets for therapeutic intervention.
Developmental Disorders: Glutamate receptors play a crucial role in brain development and neuronal circuit formation. Dysregulation of glutamate signaling during critical periods of brain development has been associated with developmental disorders, such as autism spectrum disorders (ASD). Investigating glutamate receptors in the context of developmental disorders can provide insights into their underlying mechanisms and potential therapeutic avenues.
Glutamate receptors are a topic of extensive research due to their central role in brain function and their involvement in various neurological and psychiatric conditions. Understanding the mechanisms and modulation of glutamate receptors can lead to advancements in neuroscience, pharmacology, and the development of targeted therapies.
Available Resources for Glutamate Receptors
Creative BioMart offers a variety of recombinant proteins, cell and tissue lysates, and protein pre-coupled magnetic beads that are related to glutamate receptors. These products are specifically designed to assist you in performing a range of studies such as Western blotting, proteome analysis, and protein interaction research.
We also provide customization services that include protein expression, purification, and assay development, which are tailored to address your specific needs and inquiries about contactin research.
Additionally, we offer a wide range of technical resources, including application notes and experimental protocols, to improve your understanding and increase the success of your glutamate receptors research experiments.
The following are glutamate receptors-related molecules. Click on the molecule/target to view the research reagents.
Reference:
- Tarland A E. Effects of 2-Bromoterguride, a Dopamine D 2 Receptor Partial Agonist, in Animal Models for Negative Symptoms and Cognitive Dysfunctions Associated With Schizophrenia[M]. Freie Universitaet Berlin (Germany), 2018.

