What is HRAS Protein
The official full name of HRAS, Harvey Rat Sarcoma Viral Oncogene Homolog, may sound complex, but its significance lies in its role as a member of the Ras protein family. Synonyms for HRAS include H-Ras, HRAS1, and c-H-ras, reflecting its various names in scientific literature.
Belonging to the small GTPase superfamily, HRAS is classified as a proto-oncogene, and its structure is characterized by a G domain, which binds guanine nucleotides, and a flexible C-terminal membrane anchor. Recent research advances have shed light on the dynamic nature of HRAS, revealing its involvement in cellular signaling pathways that regulate growth, differentiation, and survival.
HRAS Biological Functions and Molecular Mechanisms
HRAS is not just a mere molecular entity; it serves as a molecular switch in cellular processes, toggling between active (GTP-bound) and inactive (GDP-bound) states. The biological functions of HRAS are diverse, influencing key cellular processes such as cell proliferation, differentiation, and survival.
At the molecular level, HRAS functions by interacting with downstream effector proteins, including RAF kinases and PI3-kinase, initiating cascades that regulate gene expression and cellular responses. The molecular mechanisms involve the activation of these effectors, leading to a series of events that ultimately dictate the fate of the cell.
HRAS Related Signaling Pathway
The HRAS signaling pathway is a complex network of interactions that govern cellular responses to external stimuli. Activation of HRAS triggers a cascade of events, with RAF kinases, MEK, and ERK playing key roles in transducing signals to the nucleus. This pathway is tightly regulated to ensure precise control over cellular processes.
Dysregulation of the HRAS signaling pathway is a hallmark of many cancers, making it an attractive target for therapeutic interventions. Researchers are actively exploring novel ways to modulate HRAS activity, with the aim of developing targeted therapies that can selectively inhibit cancer cells while sparing normal cells.
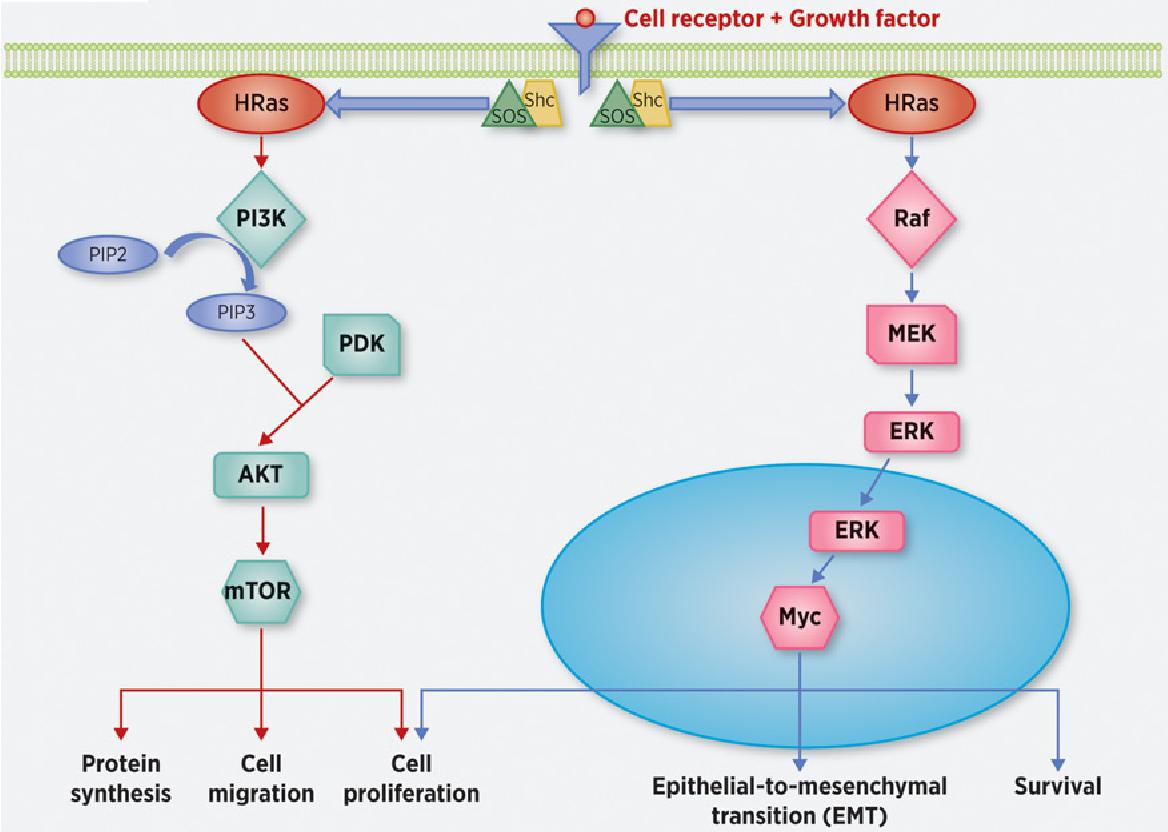
Figure 1. H-Ras signaling pathways. (Shu L, et al., 2020)
HRAS Related Diseases
While HRAS is indispensable for normal cellular functions, dysregulation of its activity can lead to pathological conditions. Mutations in HRAS have been linked to various cancers, particularly in tissues where HRAS is highly expressed. The aberrant activation of HRAS contributes to uncontrolled cell proliferation and tumorigenesis.
One well-known disorder associated with HRAS mutations is Costello syndrome, a rare genetic disorder characterized by developmental delays, intellectual disabilities, and an increased risk of tumors. Understanding the link between HRAS and diseases is crucial for developing targeted therapies and interventions.
HRAS's Applications in Biomedicine
Beyond its role in disease, HRAS has promising applications in the biomedical field. Researchers are leveraging the unique characteristics of HRAS for diagnostic development, vaccine development, and therapeutic strategies.
In diagnostic development, HRAS is being explored as a potential biomarker for certain cancers. Detecting HRAS mutations or overexpression could aid in early cancer diagnosis and prognosis, enabling timely and targeted interventions.
Vaccine development is another frontier where HRAS is making strides. Utilizing the protein's immunogenic properties, researchers are exploring the development of vaccines that target HRAS-expressing cancer cells, potentially offering a novel approach to cancer immunotherapy.
In the realm of therapeutics, targeting HRAS and its associated signaling pathways is a promising strategy for cancer treatment. Small molecule inhibitors and other targeted therapies are in development, with the goal of disrupting the abnormal signaling that drives cancer progression.
Recommended Products
| Cat.# | Product name | Species | Source (Host) | Tag |
|---|---|---|---|---|
| HRAS-2524H | Recombinant Human HRAS, His tagged | Human | Insect Cell | His |
| HRAS-2135H | Recombinant Human HRAS Protein | Human | E.coli | |
| HRAS-701H | Recombinant Human HRAS (Q61L) Protein | Human | E.coli | N/A |
| HRAS-589H | Recombinant Human HRAS, His-tagged | Human | E.coli | His |
| HRAS-229H | Recombinant Human HRAS(Mature form), His-tagged | Human | E.coli | His |
| HRAS-702H | Recombinant Human HRAS (G12V) Protein, GST-tagged | Human | E.coli | GST |
| HRAS-5023H | Recombinant Human HRAS Protein, GST-tagged | Human | Wheat Germ | GST |
| HRAS-29173TH | Recombinant Human HRAS, His-tagged | Human | E.coli | His |
| HRAS-13934H | Recombinant Human HRAS, His-tagged | Human | E.coli | His |
| HRAS-3319H | Recombinant Human HRAS Protein, Myc/DDK-tagged, C13 and N15-labeled | Human | HEK293T | Myc/DDK |
Reference
- Shu L, et al. A historic perspective and overview of H-Ras structure, oncogenicity, and targetin. Molecular Cancer Therapeutics. 2020, 19(4): 999-1007.

