Translation Regulation by the Akt Pathway
Related Symbol Search List
Immunology Background
About Translation Regulation by the Akt Pathway
The Akt pathway is a crucial signaling pathway involved in various cellular processes, including translation regulation. Translation is the process by which cellular ribosomes synthesize proteins using messenger RNA (mRNA) templates. The Akt pathway plays a significant role in controlling this process by regulating the activity of key translation factors.
One critical translation factor regulated by the Akt pathway is the mammalian target of rapamycin complex 1 (mTORC1). mTORC1 is a protein complex that senses nutrient availability and growth factor signaling to regulate protein synthesis. The Akt pathway directly phosphorylates and activates mTORC1, leading to an increase in protein synthesis.
Another translation factor regulated by the Akt pathway is eukaryotic initiation factor 4E-binding protein 1 (4E-BP1) . 4E-BP1 inhibits translation initiation by binding to eukaryotic initiation factor 4E (eIF4E), preventing its interaction with the mRNA cap structure. Akt phosphorylates and inactivates 4E-BP1, allowing eIF4E to initiate translation.
Additionally, the Akt pathway regulates the activity of the ribosomal protein S6 kinase 1 (S6K1) , another key player in translation regulation. S6K1 phosphorylates ribosomal protein S6 (rpS6) and other translation factors, promoting translation initiation and elongation.
Overall, the Akt pathway plays a crucial role in the regulation of translation by directly phosphorylating and activating multiple translation factors. This regulation allows cells to respond to nutrient availability, growth factor signaling, and other environmental cues to maintain protein synthesis homeostasis and support essential cellular functions.
Mechanism of Action of Translation Regulation by the Akt Pathway
The Akt pathway plays a crucial role in regulating several cellular processes, including cell growth, survival, and metabolism. One of the key processes controlled by the Akt pathway is translation, which is the mechanism by which genetic information is converted into functional proteins.
Translation regulation by the Akt pathway involves multiple steps, starting with the activation of Akt itself. Akt is a serine/threonine kinase that is activated by various growth factors, hormones, and other cellular signals. Upon activation, Akt phosphorylates several downstream targets, including mTORC1, TSC2, eIF4B, GSK3, and 4E-BP1.
Akt phosphorylates and activates mTORC1, which then promotes translation initiation. mTORC1 stimulates the phosphorylation of the translation repressor 4EBP1, leading to its dissociation from the eukaryotic translation initiation factor 4E (eIF4E). This allows eIF4E to associate with eIF4G and eIF4A, forming the eIF4F complex that binds to the 5' cap structure of mRNA and initiates translation.
Akt also phosphorylates and inhibits the tuberous sclerosis complex 2 (TSC2), leading to the activation of mTORC1. TSC2 normally inhibits mTORC1 activity by promoting the conversion of active GTP-bound Rheb (Ras homolog enriched in the brain) to inactive GDP-bound Rheb. Phosphorylation of TSC2 by Akt inhibits its activity, resulting in sustained activation of mTORC1 and increased translation.
In addition to mTORC1, Akt also phosphorylates and activates the eukaryotic initiation factor 4B (eIF4B). eIF4B is a translation initiation factor that enhances the helicase activity of eIF4A, promoting the unwinding of mRNA secondary structures and facilitating translation initiation.
Furthermore, Akt phosphorylates and inactivates glycogen synthase kinase 3 (GSK3). GSK3 negatively regulates translation by phosphorylating and inhibiting the initiation factor eIF2B. Akt-mediated inhibition of GSK3 relieves this inhibition and promotes translation.
Overall, the Akt pathway regulates translation by activating mTORC1, activating eIF4B, and inhibiting GSK3. These mechanisms coordinate and fine-tune protein synthesis in response to various extracellular signals to maintain cellular homeostasis.
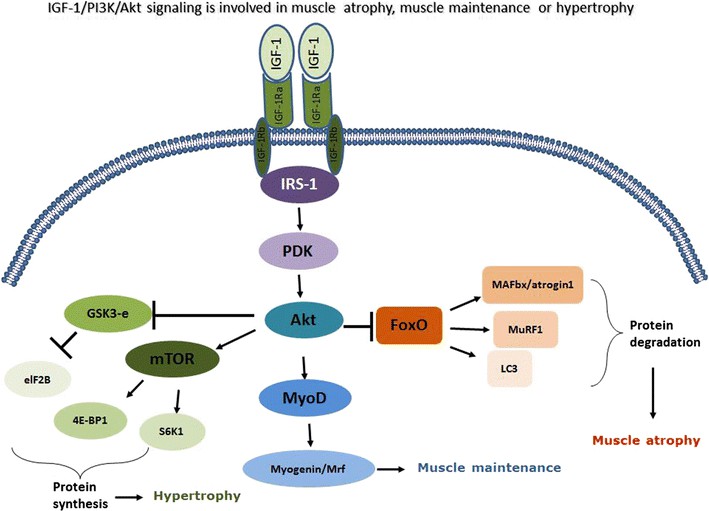 Fig.1 The IGF-1-Akt axis controls both protein synthesis and protein degradation. (Malavaki C J, et al., 2015)
Fig.1 The IGF-1-Akt axis controls both protein synthesis and protein degradation. (Malavaki C J, et al., 2015)
Available Resources for Translation Regulation by the Akt Pathway
Creative BioMart is a bioscience-focused company offering products and customizable services related to the translational regulation of the Akt pathway, as well as a wide range of other resources. Our goal is to facilitate the advancement of scientific research by providing high-quality products and specialized technical support to researchers and biopharmaceutical companies. Our products include Akt pathway-related recombinant proteins, cell and tissue lysates, and protein pre-coupled magnetic beads to meet different experimental needs.
Below are molecules/targets related to Akt pathway translation regulation, click to view product details.
We are also committed to providing a wealth of other resources, including application-specific technical guides, experimental protocols, and specialized customer support.
Through our abundant resources and professional team, we are committed to providing our customers with the highest quality solutions to help them achieve scientific success. Whether it is basic research or drug discovery, we are dedicated to providing our customers with the best support and services.
Contact us today to learn how our products and customized services can advance your research into the translational regulation of the Akt pathway.
Reference:
- Malavaki C J, Sakkas G K, Mitrou G I, et al. Skeletal muscle atrophy: disease-induced mechanisms may mask disuse atrophy[J]. Journal of muscle research and cell motility, 2015, 36: 405-421.

