Map2
-
Official Full Name
microtubule-associated protein 2 -
Overview
This gene encodes a protein that belongs to the microtubule-associated protein family. The proteins of this family are thought to be involved in microtubule assembly, which is an essential step in neurogenesis. The products of similar genes in rat and mouse are neuron-specific cytoskeletal proteins that are enriched in dentrites, implicating a role in determining and stabilizing dentritic shape during neuron development. A number of alternatively spliced variants encoding distinct isoforms have been described. [provided by RefSeq, Jan 2010] -
Synonyms
MAP2;microtubule-associated protein 2;MAP2A;MAP2B;MAP2C;MAP-2
Recombinant Proteins
- Human
- Mouse
- Rat
- HEK293
- Wheat Germ
- Mammalian Cells
- E.coli
- In Vitro Cell Free System
- Yeast
- Avi
- Fc
- His
- Non
- GST
- DDK
- Myc
- Flag
- SUMO
Background
What is MAP2 protein?
MAP2 gene (microtubule associated protein 2) is a protein coding gene which situated on the long arm of chromosome 2 at locus 2q34. This gene encodes a protein that belongs to the microtubule-associated protein family. The proteins of this family are thought to be involved in microtubule assembly, which is an essential step in neurogenesis. The products of similar genes in rat and mouse are neuron-specific cytoskeletal proteins that are enriched in dentrites, implicating a role in determining and stabilizing dentritic shape during neuron development. The MAP2 protein is consisted of 1827 amino acids and MAP2 molecular weight is approximately 199.5 kDa.
What is the function of MAP2 protein?
It is mainly expressed in the dendrites of neurons and plays an important role in the structure and stability of microtubules, neuronal morphogenesis, cytoskeleton dynamics, and organelle transport in axons and dendritic cells. By binding to microtubules, MAP2 promotes the polymerization of microtubules, forms a bundle structure, and cross-links with other cell structures. In addition, MAP2 is involved in the regulation of intracellular signaling, such as as an anchor protein for cAMP-dependent protein kinase A(PKA), It affects the amount of PKA and the phosphorylation of substrate CREB(cAMP-responsive element binding protein), thus playing a role in nerve cell signal transduction. During cell division, MAP2 is also involved in the assembly and stabilization of microtubule networks, which has an impact on the migration of nerve cells and the organizational structure of the cerebral cortex.
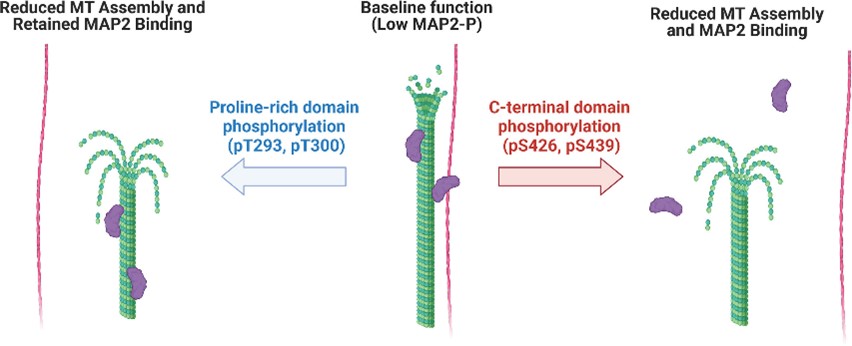
Fig1. Model of MAP2 regulation by phosphorylation in proline-rich and C-terminal domains. (R A DeGiosio, 2023)
MAP2 Related Signaling Pathway
As the anchor protein of PKA, MAP2 plays a role in nerve cell signal transduction, affecting the amount of PKA and the phosphorylation of substrate CREB. MAP2 forms the spindle of microtubules during mitosis and binds to centrioles in the middle of the spindle to participate in the formation of pseudopods and guide chromosome separation. MAP2 is involved in the intracellular transport of vesicles and particles along microtubules because some molecular motors are able to bind to the microtubules to transport materials of the cell. MAP2 is involved in the nucleation and stabilization of microtubules through the phosphorylation process, influences the transport of organelles within axons and dendrites, and may act as an anchor protein for protein kinases, which is important for signal transduction.
MAP2 Related Diseases
The MAP2 protein has been implicated in a variety of neurological diseases, especially in neurodegeneration and tumors. For example, in Alzheimer's disease (AD), abnormal phosphorylation of MAP2 is associated with the formation of neurofibrillary tangles. In addition, MAP2 expression levels change in certain types of brain tumors, such as glioma, and their expression levels correlate with the grade and aggressiveness of the tumor. Abnormal expression of MAP2 is also associated with neurodevelopmental disorders and may affect neuronal differentiation and function. In neuroblastoma, the expression and distribution of MAP2 changes with cell differentiation.
Bioapplications of MAP2
Because of its unique role in the nervous system, the MAP2 protein has a variety of applications, especially in the field of neuroscience research and clinical diagnosis. As a marker of mature neurons, MAP2 is often used to detect and identify the existence and status of neurons. In neurodevelopmental studies, the expression pattern of MAP2 contributes to understanding the process of neuronal differentiation and maturation. In pathological studies, abnormal expression or phosphorylation of MAP2 can be used as a biomarker to diagnose certain neurodegenerative diseases, such as Alzheimer's disease. In addition, MAP2 is also an important marker in brain tumor research, and the change of its expression level is helpful for tumor grading, invasive evaluation and prognosis judgment. In the field of drug development, MAP2 may become a potential target for the treatment of neurodegenerative diseases and brain tumors, and by modulating the function of MAP2 or the signaling pathways in which it interacts, it may help develop new therapeutic strategies. Related MAP2 research tools are also being actively developed, including MAP2 antibody, MAP2 staining reagents, MAP2 markers, etc.
Case Study
Case Study 1: Kohei Nishida, 2023
The physical properties of cytoskeletal microtubules have a multifaceted effect on the expression of their cellular functions. A superfamily of microtubule-associated proteins, MAP2, MAP4, and tau, promote the polymerization of microtubules, stabilize the formed microtubules, and affect the physical properties of microtubules. Here, researchers show differences in the effects of these three MAPs on the physical properties of microtubules. When microtubule-binding domain fragments of MAP2, tau, and three MAP4 isoforms were added to microtubules in vitro and observed by fluorescence microscopy, tau-bound microtubules showed a straighter morphology than the microtubules bound by MAP2 and the three MAP4 isoforms. Flexural rigidity was evaluated by the shape of the teardrop pattern formed when microtubules were placed in a hydrodynamic flow, revealing that tau-bound microtubules were the least flexible.
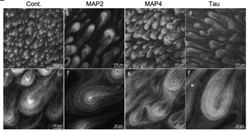

Fig1. Fluorescence microscopic images of teardrop patterns formed in the presence or absence (Cont.) of microtubule-binding domain (MBD) fragments of MAPs.
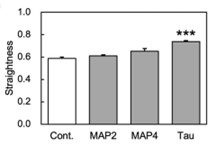
Fig2. Relationship between types of MAPs added and straightness.
Case Study 2: M Chiara Perego, 2023
Arsenic exposure during embryogenesis can lead to improper neurodevelopment and changes in locomotor activity. Additionally, in vitro studies have shown that arsenic inhibits the differentiation of sensory neurons and skeletal muscle. In the current study, human-induced pluripotent stem (iPS) cells were differentiated into motor neurons over 28 days, while being exposed to up to 0.5 μM arsenic. On day 6, neuroepithelial progenitor cells (NEPs) exposed to arsenic had reduced transcript levels of the neural progenitor/stem cell marker nestin (NES) and neuroepithelial progenitor marker SOX1, while levels of these transcripts were increased in motor neuron progenitors (MNPs) at day 12. RNA sequencing demonstrated that the cholinergic synapse pathway was impaired following exposure to 0.5 μM arsenic, and that transcript levels of genes involved in acetylcholine synthesis (CHAT), transport (solute carriers, SLC18A3 and SLC5A7) and degradation (acetylcholinesterase, ACHE) were all downregulated in day 18 early MNs. In day 28 mature motor neurons, arsenic significantly downregulated protein expression of microtubule-associated protein 2 (MAP2) and ChAT by 2.8- and 2.1-fold, respectively, concomitantly with a reduction in neurite length.
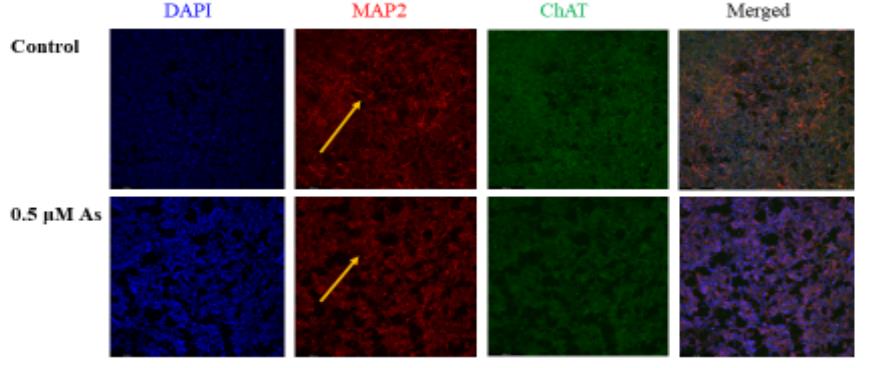
Fig3. Representative images of MAP2 (red) and ChAT (green) in control.
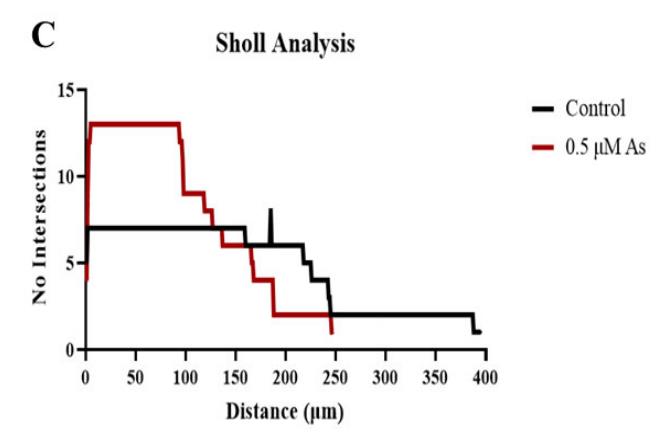
Fig4. Sholl analysis of neural processes of MAP2+ day 28 mature MNs was performed on one to two cells per biological replicate.
Quality Guarantee
High Purity
.jpg)
Fig1. SDS-PAGE (MAP2-6820H)
.
.jpg)
Fig2. SDS-PAGE (MAP2-047H)
Involved Pathway
Map2 involved in several pathways and played different roles in them. We selected most pathways Map2 participated on our site, such as LKB1 signaling events,MAPK Cascade,SIDS Susceptibility Pathways, which may be useful for your reference. Also, other proteins which involved in the same pathway with Map2 were listed below. Creative BioMart supplied nearly all the proteins listed, you can search them on our site.
| Pathway Name | Pathway Related Protein |
|---|---|
Protein Function
Map2 has several biochemical functions, for example, calmodulin binding,dystroglycan binding,microtubule binding. Some of the functions are cooperated with other proteins, some of the functions could acted by Map2 itself. We selected most functions Map2 had, and list some proteins which have the same functions with Map2. You can find most of the proteins on our site.
| Function | Related Protein |
|---|---|
Interacting Protein
Map2 has direct interactions with proteins and molecules. Those interactions were detected by several methods such as yeast two hybrid, co-IP, pull-down and so on. We selected proteins and molecules interacted with Map2 here. Most of them are supplied by our site. Hope this information will be useful for your research of Map2.
GRB2;CalpB;JUNB;MDM2;CBFB;SCHIP1;3',5'-cyclic amp;8-aha-camp;8-aha-omethyladenosine;PTBP3;p27958-pro_0000037566
Resources
Related Services
Related Products
References
- Liang, CM; Weng, SJ; et al. Neurotrophic and neuroprotective potential of human limbus-derived mesenchymal stromal cells. CYTOTHERAPY 16:1371-1383(2014).
- Cherry, JF; Bennett, NK; et al. Engineered N-cadherin and L1 biomimetic substrates concertedly promote neuronal differentiation, neurite extension and neuroprotection of human neural stem cells. ACTA BIOMATERIALIA 10:4113-4126(2014).



