NFKBIA
-
Official Full Name
nuclear factor of kappa light polypeptide gene enhancer in B-cell extracts inhibitor, alpha -
Overview
IκBα (Inhibitor of Nuclear Factor (NF)-κB α isoform) is a member of the IκB protein family with a MW of 40 kDa. IκBα is ubiquitously expressed among mammals. Located at IκBα's N-terminus is a signal response domain which contains serine residues that can be phosphorylated, while the C-terminus contains a PEST domain, a common feature among proteins with high turnover rates. As with all members of the IκB family, IκBα possesses ankyrin repeats approximately 33 amino acids in length, which mediate binding to the Rel homology region of NF-κB. The interaction of IκBα with NF-κB masks the nuclear localization sequence of NF-κB, preventing NF-κB translocation to the nucleus. A variety of stimuli can activate gene expression by liberating NF-κB through the degradation of IκBα. These stimuli include the proinflammatory cytokines TNF-α and IL-1β, chemokines, PMA, growth factors, LPS, UV irradiation, and viral infection, as well as various chemical and physical stresses. The series of events leading to this liberation is well-defined. In response to stimulus, Ser32/36 of IκBα are phosphorylated, which provides a signal for IκBα E3 ligase, a protein complex composed of SKP-1, Cul-1, Roc1, and Fbw1. IκBα E3 ligase polyubiquitinates IκBα at lysine residues 21 and 22, and the polyubiquitinated IκBα is then targeted to the 26S proteasome for degradation. Liberated NF-κB is transported across the nuclear membrane, where it activates transcription by binding to the consensus sequence GGRNNYYCC, which is found in the promoter regions of a large number of genes including IL-6, VEGF, VCAM-1, ICAM-1, HIV long terminal repeat, and many others. The initial event which targets IκBα for degradation is phosphorylation of Ser32/36. This phosphorylation is catalyzed by a protein complex known as IKK (IκB kinase). IKK contains two kinase subunits designated IKKα (MW=85 kDa) and IKKβ (MW=87 kDa), and scaffold protein designated IKKγ/NEMO (MW=48 kDa). The kinetic properties of IKK are regulated by complex formation as well as by phosphorylation events catalyzed by upstream kinases including members of the MAPK cascade, the SAPK/JNK cascade and NIK. Through its regulation of NF-κB, IκBα controls immune and inflammatory responses, cell division, and apoptosis. Numerous disease states including arthritis, asthma, and inflammatory bowel disease are associated with loss of IκBα regulation. The importance of regulation of IκBα in cancer is underscored by the observation that multiple myelomas often possess polymorphisms at IκBα regulatory sites, and certain symptoms can be ameliorated using the proteasome inhibitor PS-341 which favors the sequestration of NF-κB in the cytoplasm. -
Synonyms
IKBA;MAD-3;NFKBI;I-kappa-B-alpha;IkappaBalpha;NF-kappa-B inhibitor alpha;ikB-alpha;major histocompatibility complex enhancer-binding protein MAD3;nuclear factor of kappa light chain gene enhancer in B-cells
Recombinant Proteins
- Human
- Mouse
- Chicken
- Rhesus macaque
- Rat
- E.coli
- Mammalian Cells
- HEK293
- Rabbit
- His
- GST
- MBP
- Non
- Avi
- Fc
- DDK
- Myc
Background
What is NFKBIA protein?
NFKBIA (NFKB inhibitor alpha) gene is a protein coding gene which situated on the long arm of chromosome 14 at locus 14q13. This gene encodes a member of the NF-kappa-B inhibitor family, which contain multiple ankrin repeat domains. The encoded protein interacts with REL dimers to inhibit NF-kappa-B/REL complexes which are involved in inflammatory responses. The encoded protein moves between the cytoplasm and the nucleus via a nuclear localization signal and CRM1-mediated nuclear export. The NFKBIA protein is consisted of 317 amino acids and its molecular mass is approximately 35.6 kDa.
What is the function of NFKBIA protein?
NFKBIA protein is an important cytokine that regulates the activity of nuclear factor κB (NF-κB). NF-κB is a transcription factor that plays a key role in immune and inflammatory responses. NFKBIA protein inhibits NF-κB activity by binding to NF-κB, preventing its entry into the nucleus and activating gene transcription. In this way, NFKBIA protein can regulate a variety of biological processes, including immune response, inflammatory response, cell proliferation, and apoptosis.
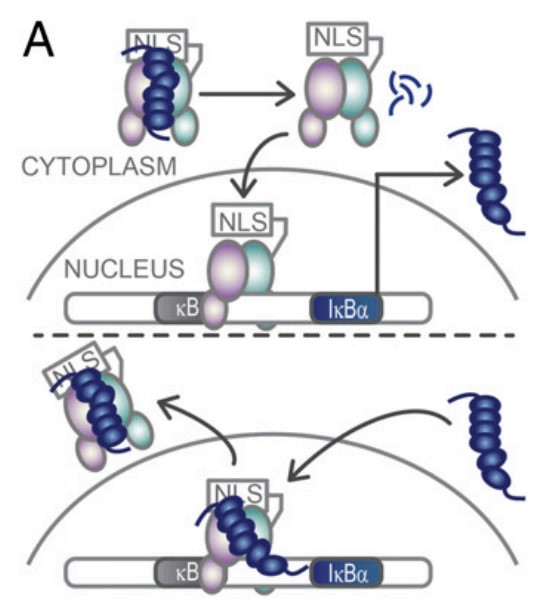
Fig1. IκBα sequesters NFκB in the cytoplasm. (Holly E Dembinski, 2017)
NFKBIA Related Signaling Pathway
NFKBIA protein is an important signaling pathway regulator, mainly involved in the nuclear factor κB (NF-κB) signaling pathway. NF-κB is a transcription factor widely found in cells and is involved in regulating many physiological and pathological processes, such as immune response, inflammation, cell growth, and apoptosis. NFKBIA proteins regulate intracellular signaling by binding to the NF-κB subunit and inhibiting its activity. In addition, NFKBIA is involved in other signaling pathways, such as tumor necrosis factor receptor (TNFR) and interleukin receptor (IL-1R), which play important roles in cellular stress, inflammation, and immune responses.
NFKBIA Related Diseases
Abnormal expression or mutation of NFKBIA has been observed in many types of cancer because NF-kB plays a key role in cell proliferation, anti-apoptosis, and inflammation. For example, breast cancer, lung cancer, colon cancer, etc. In addition, abnormal NFKBIA may also be associated with immune diseases, inflammatory diseases, and neurodegenerative diseases.
Bioapplications of NFKBIA
The application of NFKBIA protein is mainly focused on the research and treatment of diseases related to dysregulation of NF-κB activity, such as cancer, autoimmune diseases, inflammatory diseases, and neurodegenerative diseases. In addition, due to its important role in cellular stress response, NFKBIA may also have potential applications in areas such as anti-aging and tissue repair.
Case Study
Case study 1: Yanquan Li, 2023
The nuclear factor-κB (NF-κB) signaling pathway regulates specific immunological responses and controls a wide range of physiological processes. NF-κB inhibitor alpha (IKBA) is an NF-κB inhibitory mediator in the cytoplasm that modulates the nuclear translocation and DNA binding activities of NF-κB proteins. However, whether the upstream cascade of the canonical NF-κB signaling pathway has physiological roles independent of IKBA-mediated transcriptional activation remains unclear. Herein the researchers investigated the function of IKBA in mature sperm in which transcriptional and translational events do not occur. IKBA was highly expressed in human sperm. The repression of IKBA phosphorylation by its inhibitor Bay117082 markedly enhanced sperm motility. On the contrary, lipopolysaccharide-stimulated IKBA phosphorylation significantly decreased sperm motility. Nevertheless, Bay117082 treatment did not affect the motility of IKBA-knockout sperm. Further, untargeted metabolomic analysis and pharmacological blocking assays revealed that the Bay117082-induced increase in sperm motility was attributable to fatty acid β-oxidation (FAO) enhancement.
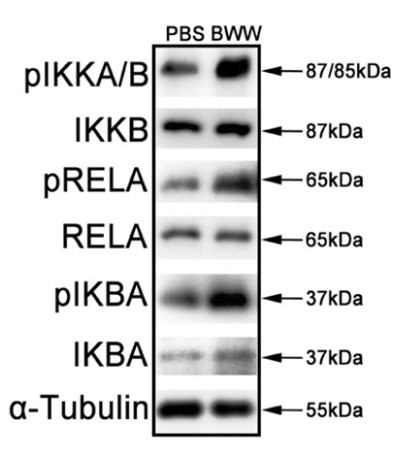
Fig1. Phosphorylation states of IKKA/B, RELA and IKBA proteins in human sperm in PBS and BWW media. α-Tubulin served as the loading control.
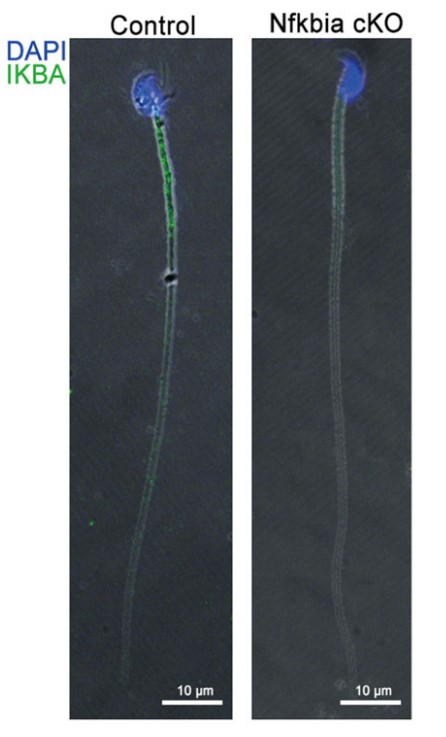
Case study 2: Holly E Dembinski, 2017
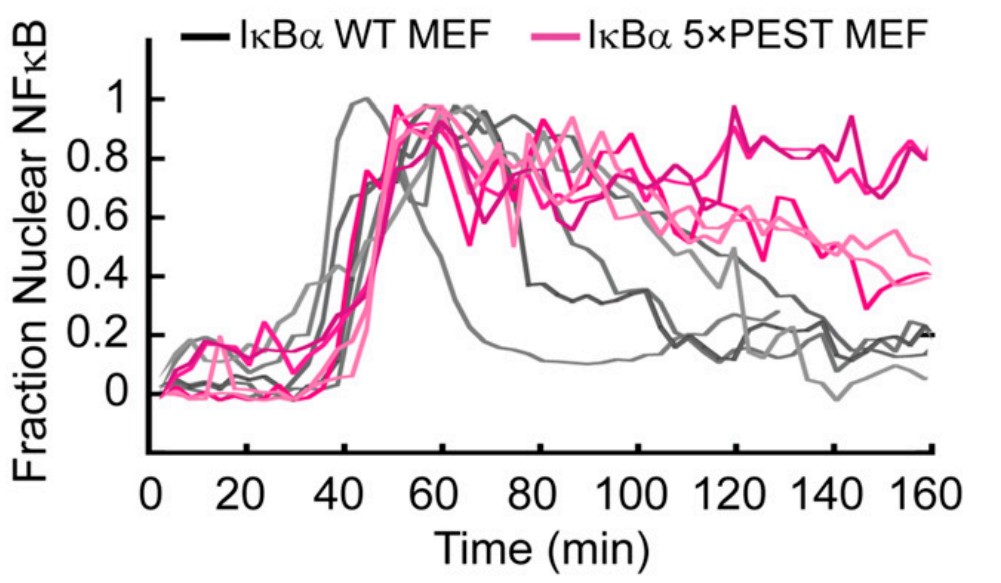
Stress-response transcription factors such as NFκB turn on hundreds of genes and must have a mechanism for rapid cessation of transcriptional activation. The researchers recently showed that the inhibitor of NFκB signaling, IκBα, dramatically accelerates the dissociation of NFκB from transcription sites, a process they have called "stripping." To test the role of the IκBα C-terminal PEST (rich in proline, glutamic acid, serine, and threonine residues) sequence in NFκB stripping, a mutant IκBα was generated in which five acidic PEST residues were mutated to their neutral analogs. This IκBα(5xPEST) mutant was impaired in stripping NFκB from DNA and formed a more stable intermediate ternary complex than that formed from IκBα(WT) because DNA dissociated more slowly. NMR and amide hydrogen-deuterium exchange mass spectrometry showed that the IκBα(5xPEST) appears to be "caught in the act of stripping" because it is not yet completely in the folded and NFκB-bound state. When the mutant was introduced into cells, the rate of postinduction IκBα-mediated export of NFκB from the nucleus decreased markedly.
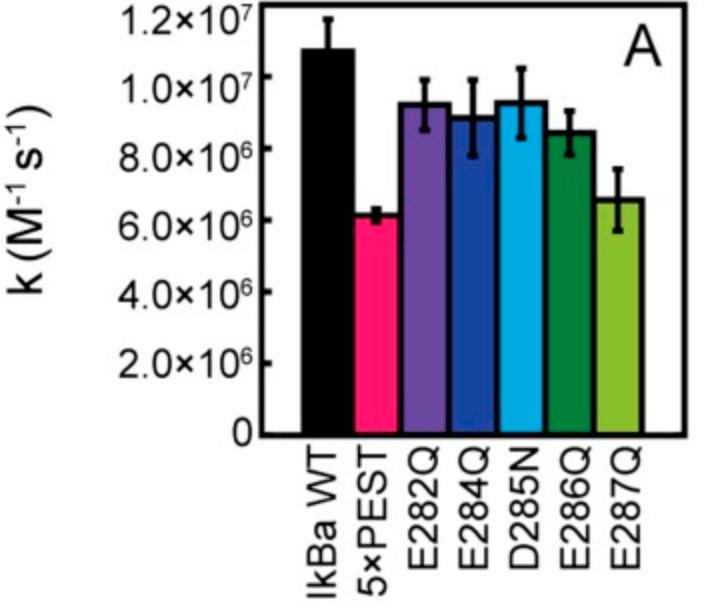
Fig3. Rates of IκBα PEST mutants associating with NFκB by stopped-flow fluorescence following the native IκBα Trp258.
Quality Guarantee
High Purity
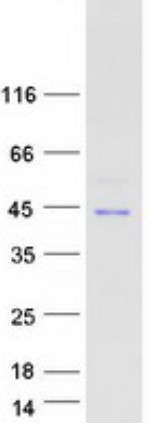
Fig1. SDS-PAGE (NFKBIA-3784H) (PROTOCOL for western blot)
High Bioactivity
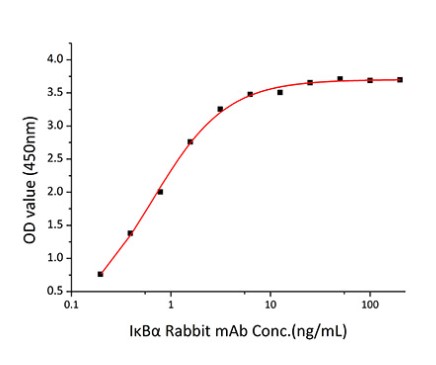
Fig2. Activity Data. (NFKBIA-2581H)
Involved Pathway
NFKBIA involved in several pathways and played different roles in them. We selected most pathways NFKBIA participated on our site, such as cAMP signaling pathway,Chemokine signaling pathway,NF-kappa B signaling pathway, which may be useful for your reference. Also, other proteins which involved in the same pathway with NFKBIA were listed below. Creative BioMart supplied nearly all the proteins listed, you can search them on our site.
| Pathway Name | Pathway Related Protein |
|---|---|
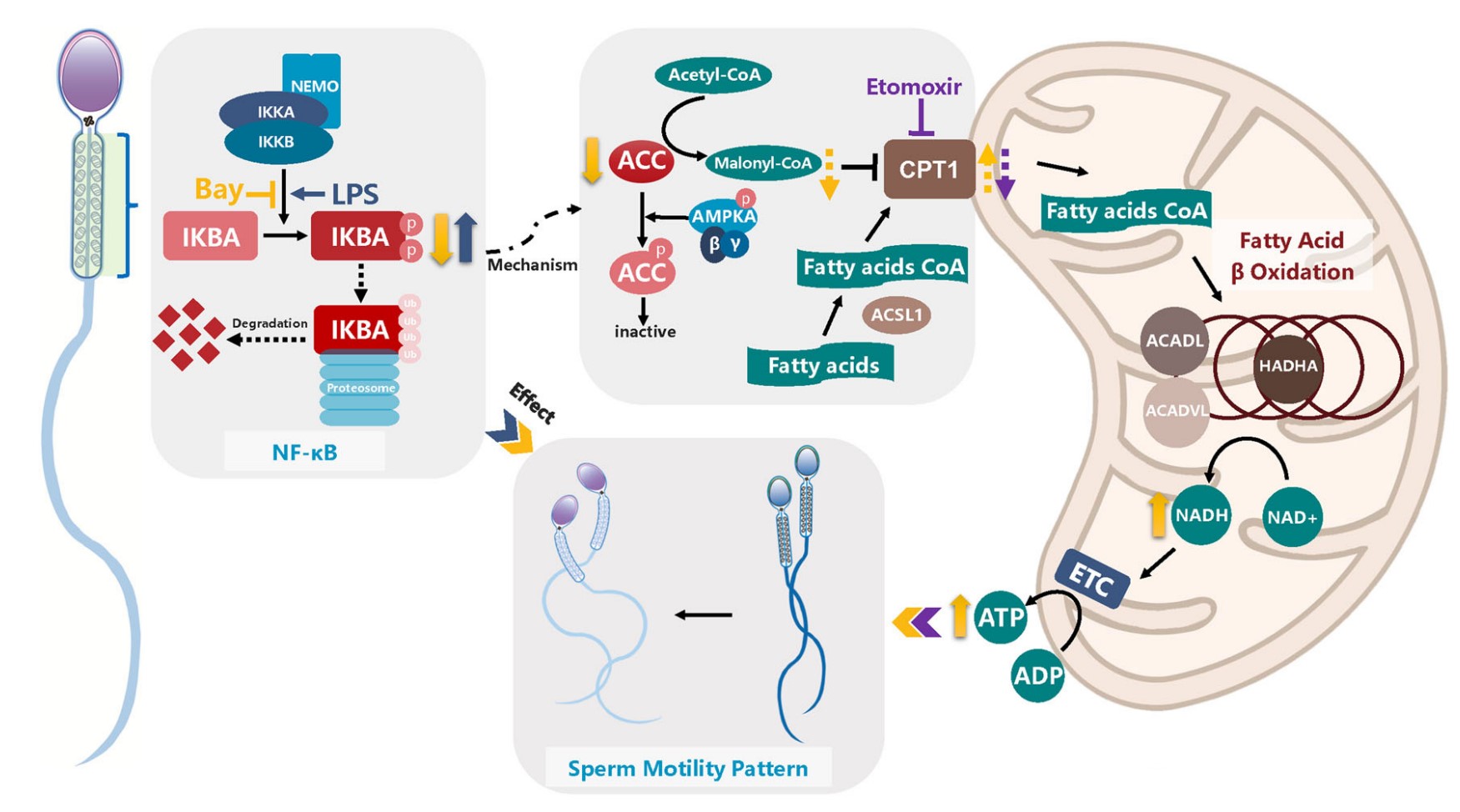
Fig1. Diagram depicting the regulation mechanism of IKBA phosphorylation in sperm motility. (Yanquan Li, 2023)
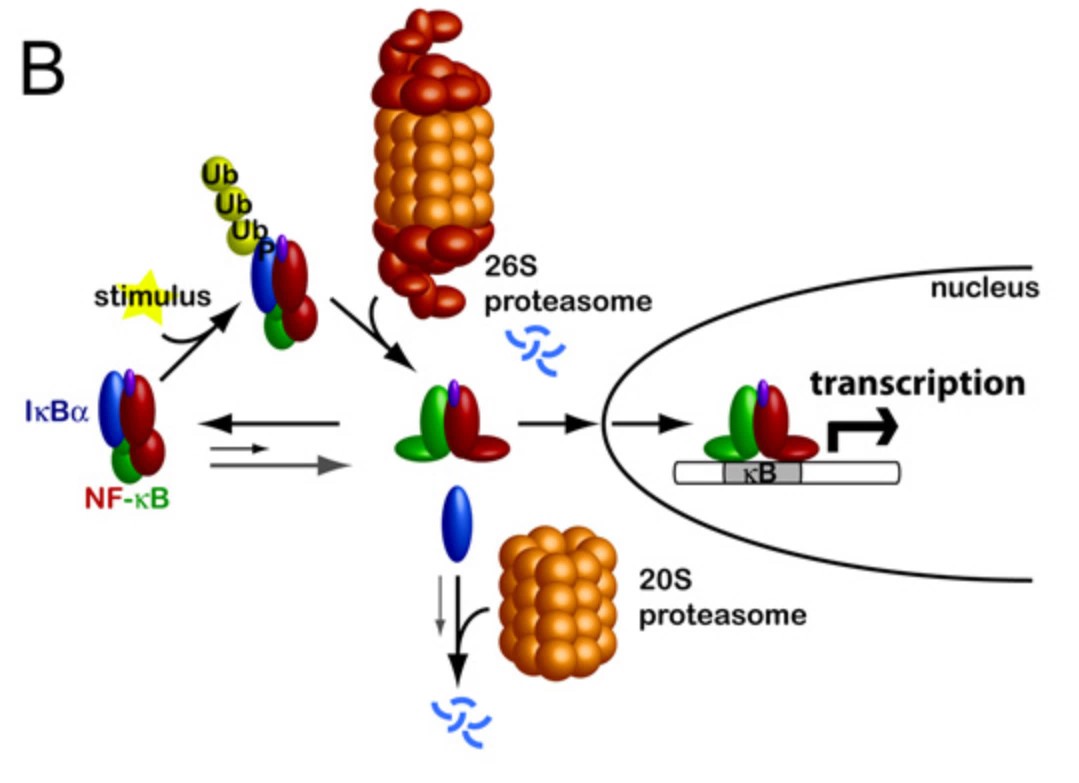
Fig2. Schematic outlining stimulus-induced activation of NF-κB. (Stephanie M E Truhlar, 2008)
Protein Function
NFKBIA has several biochemical functions, for example, NF-kappaB binding,enzyme binding,heat shock protein binding. Some of the functions are cooperated with other proteins, some of the functions could acted by NFKBIA itself. We selected most functions NFKBIA had, and list some proteins which have the same functions with NFKBIA. You can find most of the proteins on our site.
| Function | Related Protein |
|---|---|
Interacting Protein
NFKBIA has direct interactions with proteins and molecules. Those interactions were detected by several methods such as yeast two hybrid, co-IP, pull-down and so on. We selected proteins and molecules interacted with NFKBIA here. Most of them are supplied by our site. Hope this information will be useful for your research of NFKBIA.
RELA
Resources
Research Area
Related Services
Related Products
References
- Buffington, SA; Sobotzik, JM; et al. I kappa B alpha is not required for axon initial segment assembly. MOLECULAR AND CELLULAR NEUROSCIENCE 50:1-9(2012).
- Kennedy, RB; Ovsyannikova, IG; et al. Multigenic control of measles vaccine immunity mediated by polymorphisms in measles receptor, innate pathway, and cytokine genes. VACCINE 30:2159-2167(2012).



