PSMD2
-
Official Full Name
proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 -
Overview
The 26S proteasome is a multicatalytic proteinase complex with a highly ordered structure composed of 2 complexes, a 20S core and a 19S regulator. The 20S core is composed of 4 rings of 28 non-identical subunits; 2 rings are composed of 7 alpha subunits and 2 rings are composed of 7 beta subunits. The 19S regulator is composed of a base, which contains 6 ATPase subunits and 2 non-ATPase subunits, and a lid, which contains up to 10 non-ATPase subunits. Proteasomes are distributed throughout eukaryotic cells at a high concentration and cleave peptides in an ATP/ubiquitin-dependent process in a non-lysosomal pathway.An essential function of a modified proteasome, the immunoproteasome, is the processing of class I MHC peptides. This gene encodes one of the non-ATPase subunits of the 19S regulator lid. In addition to participation in proteasome function, this subunit may also participate in the TNF signalling pathway since it interacts with the tumor necrosis factor type 1 receptor. A pseudogene has been identified on chromosome 1. -
Synonyms
PSMD2;proteasome (prosome, macropain) 26S subunit, non-ATPase, 2;26S proteasome non-ATPase regulatory subunit 2;MGC14274;P97;Rpn1;S2;TRAP2;26S proteasome non ATPase regulatory subunit 2;26S proteasome regulatory subunit RPN1;26S proteasome regulatory subunit S2;26S proteasome subunit p97;55.11 protein;Proteasome (prosome macropain) 26S subunit non ATPase 2;Proteasome 26S subunit non ATPase 2;Protein 55.11;PSMD 2;PSMD2_HUMAN;TNFR associated protein 2;TRAP 2;Tumor necrosis factor receptor associated protein 2;Tumor necrosis factor type 1 receptor associated protein 2;Tumor necrosis factor type 1 receptor-associated protein 2;OTTHUMP00000210627;OTTHUMP00000210628;OTTHUMP00000210629;TNFR-associated protein 2;tumor necrosis factor receptor-associated protein 2
Recombinant Proteins
- Human
- Zebrafish
- Chicken
- Rat
- Mouse
- E.coli
- Mammalian Cells
- HEK293
- His
- Non
- Avi
- Fc
- DDK
- Myc
| Cat.# | Product name | Source (Host) | Species | Tag | Protein Length | Price |
|---|---|---|---|---|---|---|
| PSMD2-421H | Recombinant Human PSMD2 Protein, His-tagged | E.coli | Human | His | Asp645~Val872 | |
| PSMD2-10829Z | Recombinant Zebrafish PSMD2 | Mammalian Cells | Zebrafish | His |
|
|
| PSMD2-2147C | Recombinant Chicken PSMD2 | Mammalian Cells | Chicken | His |
|
|
| PSMD2-4787R | Recombinant Rat PSMD2 Protein | Mammalian Cells | Rat | His |
|
|
| PSMD2-2750HCL | Recombinant Human PSMD2 293 Cell Lysate | HEK293 | Human | Non |
|
|
| PSMD2-4446R | Recombinant Rat PSMD2 Protein, His (Fc)-Avi-tagged | HEK293 | Rat | Avi&Fc&His |
|
|
| PSMD2-4446R-B | Recombinant Rat PSMD2 Protein Pre-coupled Magnetic Beads | HEK293 | Rat |
|
||
| Psmd2-5193M | Recombinant Mouse Psmd2 Protein, Myc/DDK-tagged | HEK293 | Mouse | DDK&Myc |
|
|
| PSMD2-6044H | Recombinant Human PSMD2 Protein (Asp645-Val872), His tagged | E.coli | Human | His | Asp645-Val872 |
|
Background
What is PSMD2 protein?
PSMD2 gene (proteasome 26S subunit ubiquitin receptor, non-ATPase 2) is a protein coding gene which situated on the long arm of chromosome 3 at locus 3q27. PSMD2 protein (or known as TRAP2) is an important part of the 26S subunit of proteasome and is mainly involved in protein degradation. PSMD2 has a variety of biological functions in the cell, including apoptosis, cell cycle regulation, and cell signaling. In particular, PSMD2 acts as a key component in substrate recognition and binding in the ubiquitin-proteasome pathway, responsible for directing ubiquitin-tagged proteins to the 26S proteasome for degradation. In addition, the abnormal function of PSMD2 is associated with the occurrence and development of various diseases. The PSMD2 protein is consisted of 908 amino acids and PSMD2 molecular weight is approximately100.2 kDa.
What is the function of PSMD2 protein?
PSMD2 protein is an important part of the proteasome in the cell and is involved in the degradation of proteins. PSMD2 plays a role in a variety of biological processes, including cell cycle regulation, cell proliferation, DNA damage response, and tumor development. In tumor biology, PSMD2 is commonly used as an oncogene whose expression level is upregulated in a variety of cancers and is associated with poor clinical prognosis. For example, in lung cancer, breast cancer, and esophageal squamous cell carcinoma, high expression of PSMD2 is associated with tumor aggressiveness, lymph node metastasis, and tumor stage. PSMD2 is also involved in regulating the process of autophagy, which is critical in maintaining protein balance within cells and responding to environmental stress. In some cancers, PSMD2 promotes the survival and proliferation of tumor cells by inhibiting autophagy.
PSMD2 related signaling pathway
Psmd2-related signaling pathways are primarily involved in the ubiquitin-proteasome system (UPS), which is a major mechanism for protein degradation within cells. The protein encoded by PSMD2 is a subunit of the proteasome and is critical for its assembly, stability and its mediated protein breakdown. Ubiquitination is carried out by targeting specific proteins, i.e. attaching ubiquitin molecular markers, which are then sent to the proteasome for degradation. This process is critical for maintaining protein balance within cells, removing damaged or misfolded proteins, and regulating many cellular functions, including the cell cycle, gene expression, and cellular stress response.
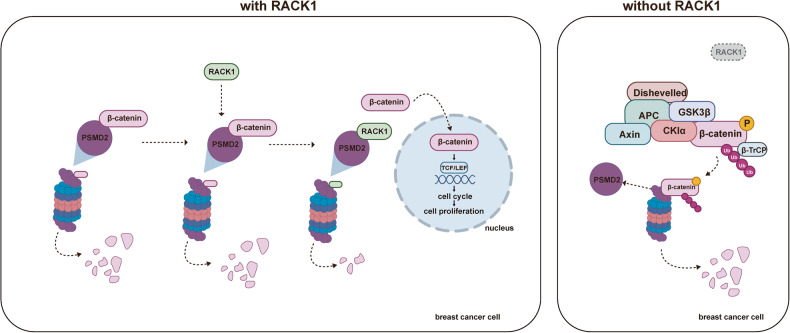
Fig1. RACK1 competitively inhibits β-catenin binding to PSMD2. (Ruinan Tian, 2023)
PSMD2 related diseases
PSMD2 protein is an important part of the proteasome in the cell and is involved in the degradation of proteins. PSMD2 is related to the occurrence and development of various diseases, especially in the field of cancer, and its abnormal expression level is closely related to the proliferation, invasion and prognosis of various tumors. For example, PSMD2 is upregulated in a variety of cancers, including non-small cell lung cancer, breast cancer, esophageal squamous cell carcinoma, and lung adenocarcinoma, and is associated with poor prognosis. In non-small cell lung cancer, PSMD2 can be used as a new therapeutic target, and its inhibition can significantly inhibit the growth and metastasis of tumor cells. In breast cancer, PSMD2 promotes cell cycle progression and cell proliferation by regulating proteasome degradation of p21 and p27. In addition, PSMD2 is also associated with levels of immune cell infiltration in the tumor immune microenvironment, which may serve as a potential target for immunotherapy.
Bioapplications of PSMD2
PSMD2 is a laboratory recombinant form of the 26S proteasome non-atpase-regulating subunit 2, which plays a key role in intracellular protein degradation. In the field of cancer therapy, rhPSMD2 may serve as a research tool to understand its mechanism of action in tumor cells, including how it regulates the degradation of cyclins such as p21 and p27, and how it contributes to the progression of certain cancer types by influencing the process of autophagy. In addition, PSMD2 expression levels are associated with poor prognosis in a variety of cancers, making it a potential biomarker and therapeutic target. Therefore, the application of rhPSMD2 in biomedical research includes the validation of drug targets, the study of disease mechanisms, and the development of novel therapeutic strategies.
Case Study
Case Study 1: Qun Zhang, 2023
Recent studies suggest that DNA methylation significantly influences the development of nasopharyngeal carcinoma (NPC). We found that DNAJA4, which is hypermethylated in NPC, is downregulated and its overexpression can inhibit NPC cell migration, invasion, and epithelial-mesenchymal transition (EMT) without affecting cell viability or proliferation. DNAJA4 promotes the degradation of MYH9 protein through the ubiquitin-proteasome pathway, and high MYH9 levels can counteract DNAJA4's suppressive effects. Clinically, low DNAJA4 levels are associated with poorer prognosis and a higher risk of distant metastasis in NPC patients.
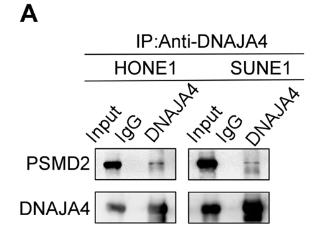
Fig1. The endogenous association of DNAJA4 and PSMD2, as well as MYH9 and PSMD2, in HONE1 and SUNE1 cells.
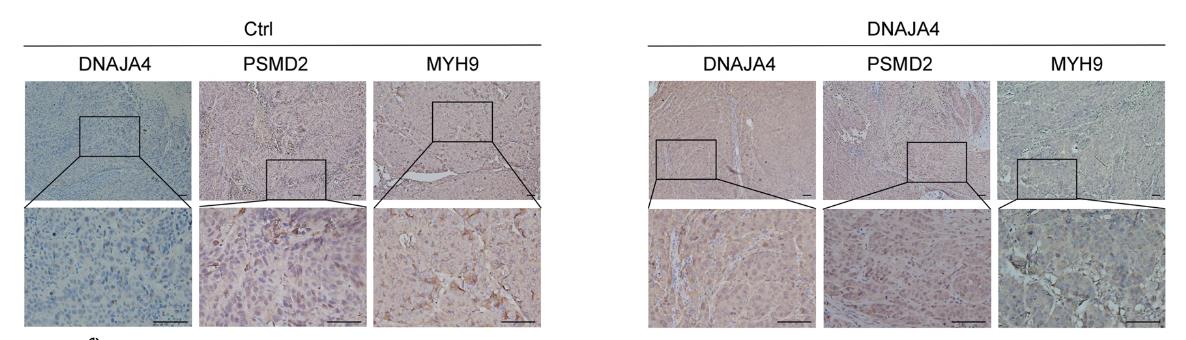
Fig2. Representative images and quantitative analysis of IHC staining of DNAJA4, PSMD2, and MYH9.
Case Study 2: Yanjie Tan, 2019
Obesity and nonalcoholic steatohepatitis (NASH) increase the risk of hepatocellular carcinoma (HCC), with a lipid-rich environment promoting tumor growth and spread. While the ubiquitin-proteasome system (UPS) is known to influence tumor cell proliferation, its role in lipid-rich tumors is not fully understood. This study found that two proteasome subunits, PSMD1 and PSMD2, control HepG2 cell proliferation by affecting lipid metabolism. Knocking down these subunits reduces lipid droplet formation, which provides energy and membrane components for tumor cells. PSMD1 and PSMD2 regulate lipid synthesis genes through p38-JNK and AKT pathways. High levels of these subunits also correlate with a worse HCC prognosis.
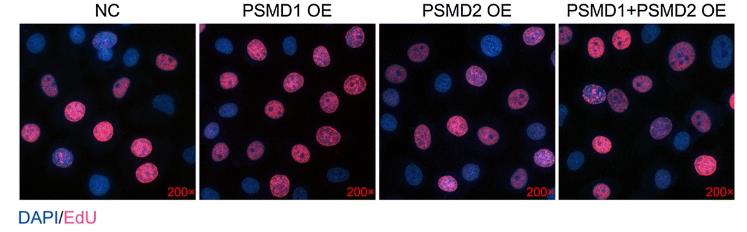
Fig3. EdU staining after PSMD1/PSMD2 overexpression.
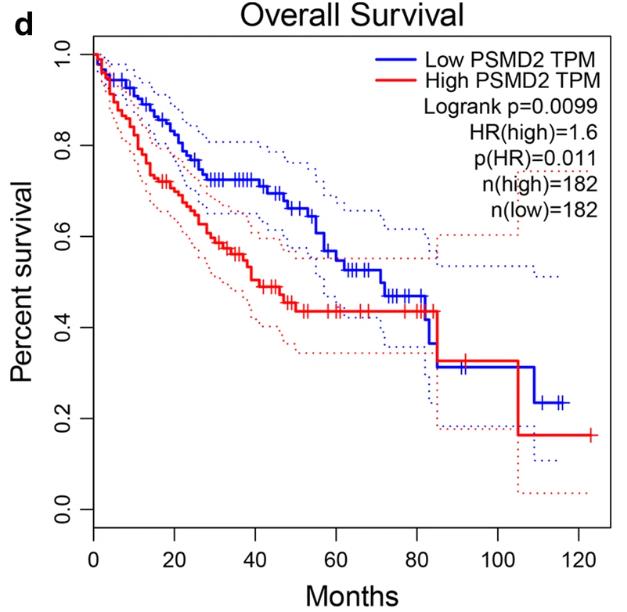
Fig4. The survival analysis of PSMD2 in LIHC.
Involved Pathway
PSMD2 involved in several pathways and played different roles in them. We selected most pathways PSMD2 participated on our site, such as Proteasome,Epstein-Barr virus infection, which may be useful for your reference. Also, other proteins which involved in the same pathway with PSMD2 were listed below. Creative BioMart supplied nearly all the proteins listed, you can search them on our site.
| Pathway Name | Pathway Related Protein |
|---|---|
| Epstein-Barr virus infection | PIK3R3,EP300,CSNK2A2,TBPL1,TNFAIP3,RBPJ,PSMC3,YWHAZ,PIK3CA,GTF2E2 |
| Proteasome | PSMA7,PSMD11,PSMB8,PSMA6A,PSMA6,PSMB9B,PSMB8F,PSMD13,PSMB4,PSMD6 |
Protein Function
PSMD2 has several biochemical functions, for example, contributes_to endopeptidase activity,enzyme regulator activity,protein binding. Some of the functions are cooperated with other proteins, some of the functions could acted by PSMD2 itself. We selected most functions PSMD2 had, and list some proteins which have the same functions with PSMD2. You can find most of the proteins on our site.
| Function | Related Protein |
|---|---|
| protein binding | BOC,ARHGEF3,EHMT2,RNASE1,ROCK2A,LRRC33,PPARA,OSBP,SUPT6H,ISYNA1 |
| enzyme regulator activity | CPN2,ATP6V1H,MTMR9,APOC3,PSMD1,RGN,BRCC3,DCP1B,GCLM,PSMD3 |
| contributes_to endopeptidase activity | PSMD1 |
Interacting Protein
PSMD2 has direct interactions with proteins and molecules. Those interactions were detected by several methods such as yeast two hybrid, co-IP, pull-down and so on. We selected proteins and molecules interacted with PSMD2 here. Most of them are supplied by our site. Hope this information will be useful for your research of PSMD2.
PSMC1;UCHL5;BAG1
Resources
Related Services
Related Products
References



