SIRT6
-
Official Full Name
sirtuin 6 -
Overview
The Silent Information Regulator (SIR2) family of genes is a highly conserved group of genes that encode nicotinamide adenine dinucleotide (NAD)-dependent protein deacetylases, also known as Class III histone deacetylases. The first discovered and best ch -
Synonyms
SIRT6;sirtuin 6;sirtuin (silent mating type information regulation 2 homolog) 6 (S. cerevisiae) , sirtuin (silent mating type information regulation 2, S. cerevisiae, homolog) 6;NAD-dependent deacetylase sirtuin-6;sirtuin type 6;SIR2-like protein 6
Recombinant Proteins
- Human
- Mouse
- Rat
- Zebrafish
- Chicken
- E.coli
- Mammalian Cells
- Sf9 Cells
- HEK293
- GST
- His
- T7
- Non
- DDK
- Myc
Background
What is SIRT6 protein?
SIRT6 (sirtuin 6) gene is a protein coding gene which situated on the short arm of chromosome 19 at locus 19p13. SIRT6 is a member of the sirtuin family of proteins which are homologs to the yeast Sir2 protein. Sirtuin family contain a sirtuin core domain and are grouped into four classes with SIRT6 being a member of class IV. Human SIRT6 protein is a NAD (+)-dependent histone H3 lysine-9 deacetylase that modulates telomeric chromatin. The SIRT6 protein is consisted of 355 amino acids and its molecular mass is approximately 39.1 kDa.
What is the function of SIRT6 protein?
SIRT6 protein is a nicotinamide adenine dinucleotide (NAD+) dependent deacetylase, which is involved in many biological processes. The main functions of SIRT6 in cells include regulation of metabolism, anti-oxidative stress, anti-inflammation and anti-apoptosis. SIRT6 is involved in the regulation of glucose, lipid and amino acid metabolism. SIRT6 protects cells from oxidative stress by regulating the expression and activity of antioxidant enzymes. SIRT6 inhibits inflammatory response and reduces the occurrence and development of inflammatory diseases. SIRT6 can inhibit apoptosis and maintain tissue homeostasis by regulating the expression of apoptosis-related genes.
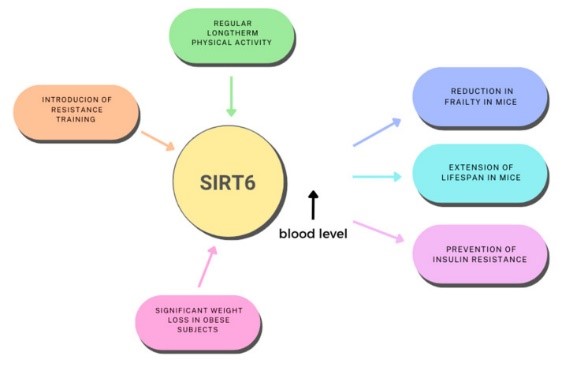
Fig1. Modifying role of physical activity and the influence of SIRT6 on metabolism. (Adrianna Dzidek, 2023)
SIRT6 Related Signaling Pathway
As an important epigenetic regulator, SIRT6 protein is involved in regulating many signaling pathways. Including: NF-κB signaling pathway, TGF-β signaling pathway, AMPK signaling pathway and so on. Among them, SIRT6 affects the activity of Wnt signaling pathway through deacetylation of β-catenin, and then regulates cell proliferation, differentiation and apoptosis. In addition, SIRT6 can also regulate AMPK, IGF-1/mTORC1, p53 and other signaling molecules, thereby regulating related signaling pathways.
SIRT6 Related Diseases
1. Neurodegenerative diseases: SIRT6 plays a protective role in neurodegenerative diseases, such as Alzheimer's disease, Parkinson's disease and Huntington's disease.
2. Tumor: SIRT6 plays a dual role in tumor genesis and development. On the one hand, SIRT6 plays an anti-tumor role by inhibiting the proliferation, invasion and metastasis of tumor cells. SIRT6, on the other hand, promotes tumor growth and drug resistance by regulating tumor cell metabolism and sensitivity to chemotherapy drugs.
3. Cardiovascular diseases: SIRT6 affects the occurrence and development of cardiovascular diseases by regulating the energy metabolism of cardiomyocytes, anti-oxidative stress and autophagy. Other related diseases include diabetes, inflammatory diseases and metabolic diseases.
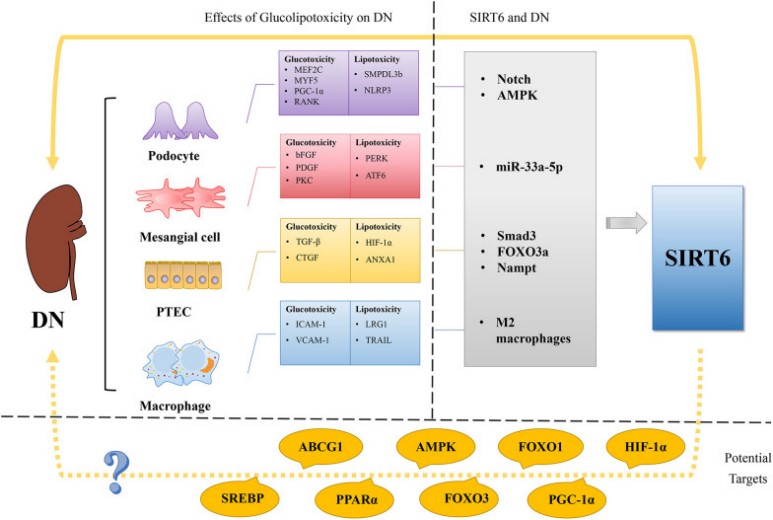
Fig2. Regulation of glucose and lipid metabolism in DN and SIRT6's possible role as a treatment target for diabetic nephropathy (DN). (Ying Wang, 2023)
Bioapplications of SIRT6
SIRT6 plays an important role in cell aging and life span regulation. Researchers are exploring ways to delay the aging process and improve quality of life and longevity by activating SIRT6. In addition, given its role in a number of diseases including neurodegenerative diseases, tumors, cardiovascular diseases, it is also being explored as a therapeutic target for drug therapy.
Case Study
Case study 1: Cristina Andreani, 2023
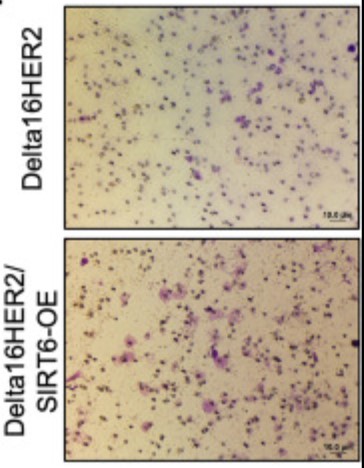
The histone deacetylase sirtuin 6 (SIRT6) has been endowed with anti-cancer capabilities in many tumor types. Here, the researchers investigate the impact of SIRT6-overexpression (SIRT6-OE) in Delta16HER2 mice, which are a bona fide model of HER2-positive breast cancer. ChIP-seq, RNA-seq and RT-PCR experiments indicate that SIRT6-OE represses the expression of the T-box transcription factor 3 (Tbx3) by deacetylation of H3K9ac.
This work indicates that high levels of SIRT6 are indicative of poor prognosis and high risk of metastasis in HER2-positive breast cancer and suggests further investigation of TBX3 as a downstream target of SIRT6 and co-marker of poor-prognosis. The results point to a breast cancer subtype-specific effect of SIRT6 and warrant future studies dissecting the mechanisms of SIRT6 regulation in different breast cancer subtypes.
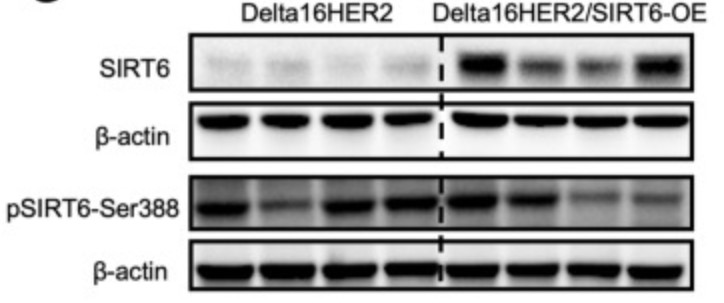
Fig1. Representative images of IHC staining for SIRT6 (brown) in tumors of 30-week-old Delta16HER2 and Delta16HER2/SIRT6-OE mice, respectively.
Case study 2: Michaela E Copp, 2023
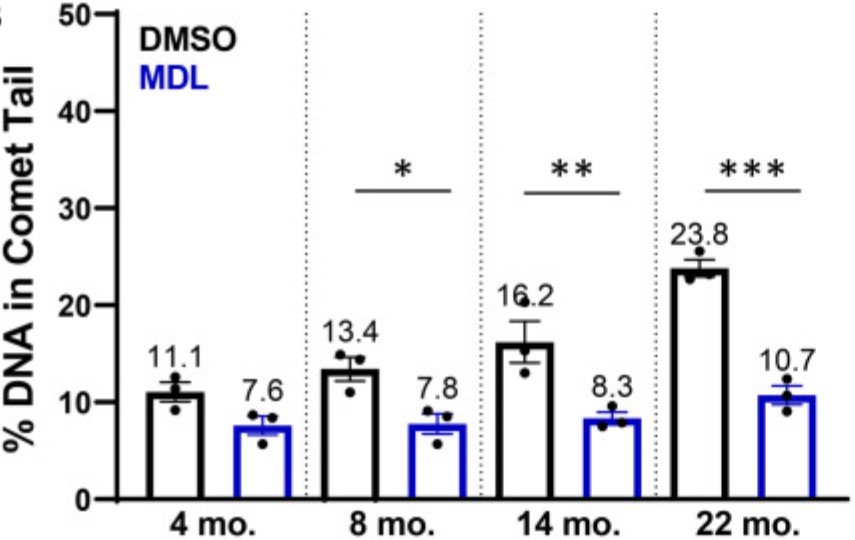
While advanced age is widely recognized as the greatest risk factor for osteoarthritis (OA), the biological mechanisms behind this connection remain unclear. The purpose of this study was to determine whether a decline in DNA repair efficiency is one explanation for the accumulation of DNA damage with age, and to quantify the improvement in repair with activation of Sirtuin 6 (SIRT6). After acute damage with irradiation, DNA repair was shown to be more efficient in chondrocytes from young (≤45 years old) as compared to middle-aged (50-65 years old) or older (>70 years old) cadaveric donors. Activation of SIRT6 with MDL-800 improved the repair efficiency, while inhibition with EX-527 reduced the rate of repair and increased the percentage of cells that retain high levels of damage.
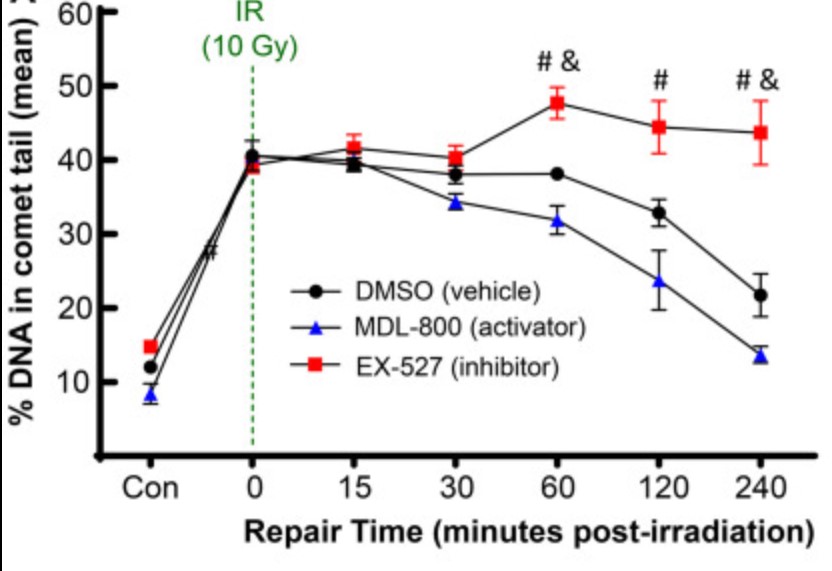
Fig3. Effect of SIRT6 activation and inhibition on chondrocyte repair rate.
Quality Guarantee
High Purity
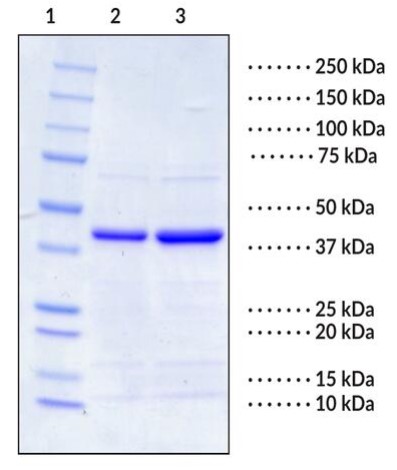
Fig1. SDS-PAGE (SIRT6-121H) (PROTOCOL for western blot)
.
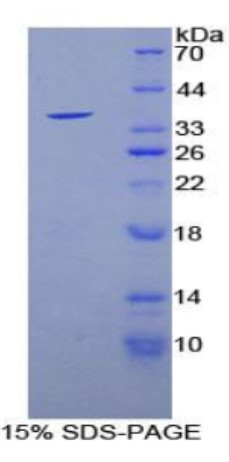
Fig2. SDS-PAGE (Sirt6-1381M) (PROTOCOL for western blot)
Involved Pathway
SIRT6 involved in several pathways and played different roles in them. We selected most pathways SIRT6 participated on our site, such as Central carbon metabolism in cancer, which may be useful for your reference. Also, other proteins which involved in the same pathway with SIRT6 were listed below. Creative BioMart supplied nearly all the proteins listed, you can search them on our site.
| Pathway Name | Pathway Related Protein |
|---|---|
| Central carbon metabolism in cancer | NRAS,PKM,PIK3CB,SCO2,PDGFRA,PDK1,KRAS,PFKM,GLS,PIK3R2 |
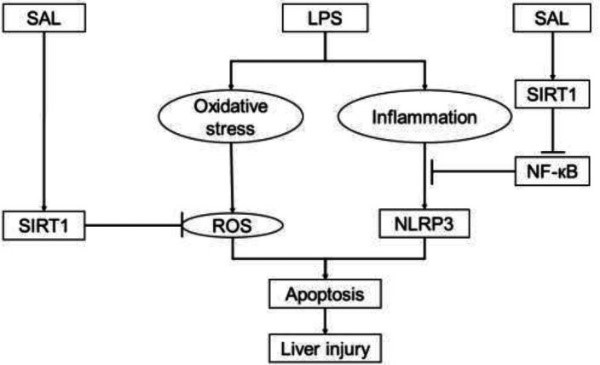
Fig1. Model illustration on SAL-mediated regulation of SIRT1/NF-κB and NLRP3 pathways, which underlies the anti-inflammatory, anti-oxidant, anti-apoptotic, and anti-liver damage actions of LPS. (Jialei Meng, 2024)
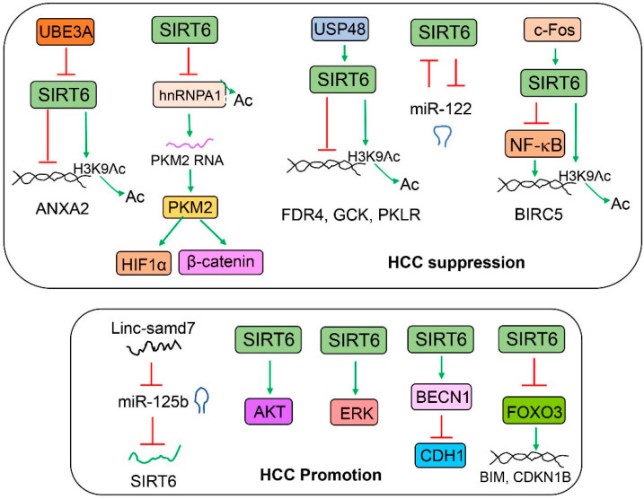
Fig2. The role of SIRT6 in liver cancer. (X Charlie Dong, 2023)
Protein Function
SIRT6 has several biochemical functions, for example, NAD(P)+-protein-arginine ADP-ribosyltransferase activity,NAD+ ADP-ribosyltransferase activity,NAD+ binding. Some of the functions are cooperated with other proteins, some of the functions could acted by SIRT6 itself. We selected most functions SIRT6 had, and list some proteins which have the same functions with SIRT6. You can find most of the proteins on our site.
| Function | Related Protein |
|---|---|
| NAD+ binding | SIRT2,SIRT7,GLUD1,SIRT3,UXS1,HADH,SIRT5,HPGD,SIRT4,ALDH1A3 |
| chromatin binding | MSGN1,PBRM1,POU1F1,TRP63,JUB,SAFB,PRDM13,TAL1,HINFP,BARX2 |
| NAD-dependent histone deacetylase activity | SIRT1,SIRT2 |
| transcription corepressor activity | TBL1XR1,HDAC7,CBX4,PSMC3,ZEB1,CIR1,TOB1,SNW1,PBXIP1,CBFA2T2 |
| NAD+ ADP-ribosyltransferase activity | PARP1,TNKS,PARP4,PARP14,ART5,GIG2P,CHAT1,TIPARP,PARP2,GIG2L |
| NAD-dependent histone deacetylase activity (H3-K9 specific) | SIRT1 |
| NAD(P)+-protein-arginine ADP-ribosyltransferase activity | ART1,ART3,CHAT1,MADPRT,ART5,CHAT2,Art2b |
| zinc ion binding | RP9,S100A8,ENPP2,ZCCHC10,USP20,MICALL2B,BHMT2,FTR82,KDM1B,RNF121 |
| protein binding | NR1I2,CDK4,NECAB3,SGTB,SPPL2B,RBBP4,POLI,HELT,EREG,COL8A1 |
Interacting Protein
SIRT6 has direct interactions with proteins and molecules. Those interactions were detected by several methods such as yeast two hybrid, co-IP, pull-down and so on. We selected proteins and molecules interacted with SIRT6 here. Most of them are supplied by our site. Hope this information will be useful for your research of SIRT6.
RELA;VIM;CHD3;MYC;a8k1f4_human;MLF1;HSF2;PSMD2;q96be0_human;AARSD1;FKBPL;STUB1;UBE2D1
Resources
Research Area
Related Services
Related Products
References
- Tahir, MN; Ragg, R; et al. Amine functionalized ZrO2 nanoparticles as biocompatible and luminescent probes for ligand specific cellular imaging. JOURNAL OF MATERIALS CHEMISTRY B 3:2371-2377(2015).
- Orellana, ME; Quezada, C; et al. Expression of SIRT2 and SIRT6 in Retinoblastoma. OPHTHALMIC RESEARCH 53:100-108(2015).



