Ovarian Cancer Biomarkers
Related Symbol Search List
Immunology Background
What Is reproductive system cancer
Reproductive cancers are cancers that occur in the reproductive organs. In women, these are cancers in the breast, cervix, uterus, vulva, endometrium or ovaries. In men, these are cancers in testicular, penile and prostate gland. Breast cancer remains the second most frequently occurred cancer worldwide as well as the most common cancer in women. Cervical cancer is a worldwide medical problem with a very disproportionate global distribution. A high incidence is seen in low resource countries, especially in Africa, Latin America, parts of Asia and in some eastern European countries with incidences up to 100/100000. In contrast, in industrialized countries, the incidence may be as low as 10/100000 women. Prostate cancer is one of the most common cancers among men and a major challenge for clinicians.
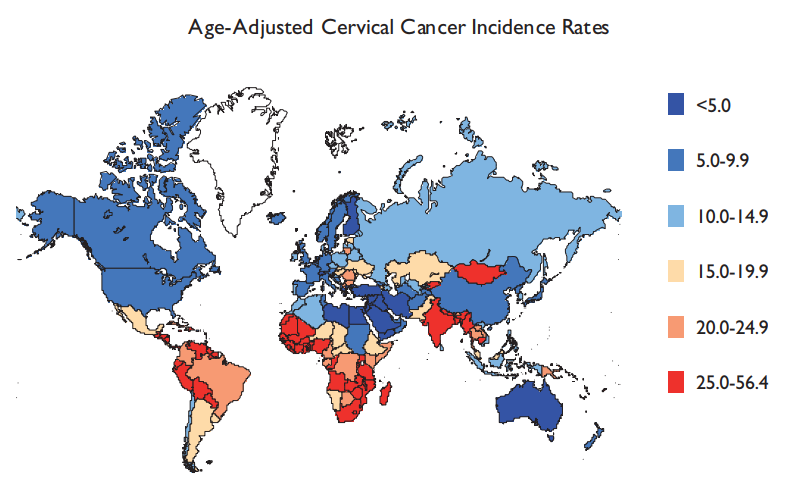 Figure 1. International Variations in Age-Adjusted Cervical Cancer Incidence Per 100,000.
Figure 1. International Variations in Age-Adjusted Cervical Cancer Incidence Per 100,000.
Source: WHO, International Agency for Research on Cancer, GLOBOCAN, 2008.
Types of reproductive system cancer
Reproductive system cancer could be classified into female reproductive cancer and male reproductive cancer. The most common reproductive cancers in women are: cervical cancer, ovarian cancer, uterine cancer, vaginal cancer and vulvar cancer. The most common reproductive cancers in men are: testicular cancer, penile cancer and prostate cancer.
Risk factors for reproductive system cancer
Table 1. Risk factors for female reproductive system cancer
| Cancer types | Risk factors |
| Breast |
Being 55 years old or older. Someone in your family having had breast cancer. Smoking. Drinking alcohol. |
| Cervical |
Smoking. Having HIV or a weak immune system. Persistence of high-risk human papillomavirus (HPV). |
| Ovarian |
Being 40 years or older. Someone in your family having had cancer of the ovary, breast, or colon. Being of Eastern European Jewish descent. Never giving birth. |
| Uterine |
Being 50 years or older. Being overweight. Someone in your family having had cancer of the uterus, ovary, or colon. |
| Vaginal & Vulvar |
Having HPV. Having cervical pre-cancer or cancer. Having HIV or a weak immune system. Smoking. |
Table 2. Risk factors for male reproductive system cancer
| Cancer types | Risk factors |
| Testicular |
Undescended testicle. Having a family history of testicular cancer. Having a testicle that is not normal. |
| Penile |
Having human papillomavirus (HPV). Being uncircumcised Being age 60 or older. Having many sexual partners, using tobacco products. |
| Prostate |
Being aged 50 years old or older. Having a family history of prostate cancer. |
Reproductive system cancer proteins
Integration of powerful prognostic biomarkers in the pathology workflow would help identify patients with an increased risk of developing an aggressive disease. Up to now, for prostate cancer patient, NF-kappa B, vimentin, prostate and breast overexpressed 1 (PBOV1) are prognostic factors. The accuracy of prostate cancer stratification would gain from the addition of new prognostic biomarkers.
For breast cancer, until now, three key protein biomarkers have shown great help in guiding prognosis and therapy, including progesterone receptor (PR), estrogen receptor (ER), and human epidermal growth factor (EGF) receptor 2 (HER2).
Tumor vascular markers (TVMs) is potential biomarkers and molecular targets for ovarian cancer and a variety of other solid tumors. Immunohistochemistry demonstrated expression of F2RL1 in a periendothelial location and in stromal cells in tumors, whereas staining in normal ovary was rare or absent. DR6 protein was diffusely expressed in tumor vascular structures and faintly in tumor stroma. DR6 protein was expressed in the vasculature of normal ovaries.
In cervical cancer, currently, a number of potential biomarkers for cervical screening are being analyzed. One of them is 28-kDaGolgiSNAREprotein (GS28).
In testicular cancer, a large number of biomarkers have been proposed for the diagnosis. The conventional tumor markers (a-fetoprotein) AFP, hCG, and LDH have demonstrated value in the clinical management of testicular malignant TGCT.
These biomarkers could have true clinical applications in the near future, mainly being used as prognostic markers to guide the aggressiveness of initial treatment with increased usage of multimodal management.
Table 3. Protein biomarkers for the detection of some types of reproductive system cancer
| Types of reproductive system cancer | Biomarkers |
| Prostate Cancer | NF-kappa B, vimentin, prostate and breast overexpressed 1 (PBOV1) |
| Breast Cancer | Progesterone receptor (PR), Estrogen receptor(ER), HER2 |
| Cervical Cancer | 28-kDaGolgiSNAREprotein (GS28) |
| Penile Carcinoma | IGFBP2 |
| Testicular Cancer | AFP, hCG, LDH |
References:
1. Grosset A A, Ouellet V, Caron C, et al. Validation of the prognostic value of NF-κB p65 in prostate cancer: A retrospective study using a large multi-institutional cohort of the Canadian Prostate Cancer Biomarker Network[J]. PLoS medicine, 2019, 16(7).
2. Jiang C, Wu S, Jiang L, et al. Network-based approach to identify biomarkers predicting response and prognosis for HER2-negative breast cancer treatment with taxane-anthracycline neoadjuvant chemotherapy[J]. PeerJ, 2019, 7: e7515.
3. Milardi D, Grande G, Vincenzoni F, et al. Proteomics for the identification of biomarkers in testicular cancer–review[J]. Frontiers in endocrinology, 2019, 10: 462.

