SOD2
-
Official Full Name
superoxide dismutase 2, mitochondrial -
Overview
This gene is a member of the iron/manganese superoxide dismutase family. It encodes a mitochondrial protein that forms a homotetramer and binds one manganese ion per subunit. This protein binds to the superoxide byproducts of oxidative phosphorylation and converts them to hydrogen peroxide and diatomic oxygen. Mutations in this gene have been associated with idiopathic cardiomyopathy (IDC), premature aging, sporadic motor neuron disease, and cancer. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. [provided by RefSeq, Jul 2008] -
Synonyms
SOD2;superoxide dismutase 2, mitochondrial;IPOB;MNSOD;MVCD6;superoxide dismutase [Mn], mitochondrial;indophenoloxidase B;Mn superoxide dismutase;mangano-superoxide dismutase;manganese-containing superoxide dismutase
Recombinant Proteins
- Human
- Zebrafish
- Rat
- Mouse
- Rhesus macaque
- Chicken
- Cynomolgus
- Macaca Fascicularis
- E.coli
- Mammalian Cells
- HEK293
- HeLa
- Non
- His
- GST
- MBP
- DDK
- Myc
- Flag
- Avi
- Fc
Background
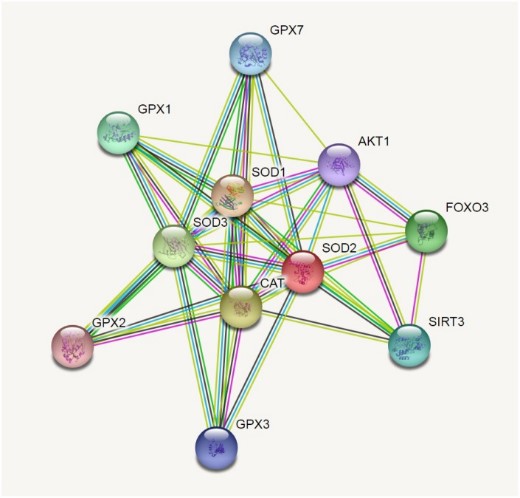
Fig1. SOD2 correlations crosstalk investigation by String server functional protein association networks (Homo sapiens).
What is SOD2 protein?
SOD2 (superoxide dismutase 2) gene is a protein coding gene which situated on the long arm of chromosome 6 at locus 6q25. The encoded protein is a member of the iron/manganese superoxide dismutase family. It encodes a mitochondrial protein that forms a homotetramer and binds one manganese ion per subunit. This protein binds to the superoxide byproducts of oxidative phosphorylation and converts them to hydrogen peroxide and diatomic oxygen. The SOD2 protein is consisted of 222 amino acids and its molecular mass is approximately 27.8 kDa.
What is the function of SOD2 protein?
SOD2 can catalyze the reaction of superoxide anion free radicals with water to generate hydroperoxide and oxygen, thereby reducing the concentration of superoxide free radicals in cells and alleviating oxidative stress damage. By scavenging superoxide free radicals produced within mitochondria, SOD2 protects mitochondrial membranes, proteins, and DNA from oxidative damage. SOD2 also reduces inflammation. Normal SOD2 activity can maintain cell stability and inhibit cell mutation and malignant transformation.
SOD2 Related Signaling Pathway
The signaling pathway related to SOD2 protein involves many aspects, including ROS/ oxidative stress signaling pathway, PI3K/AKT signaling pathway, MAPK signaling pathway, Nrf2/ARE signaling pathway and NF-κB signaling pathway. These signaling pathways interact to jointly regulate the expression and activity of SOD2 and influence the cell response to oxidative stress.
SOD2 Related Diseases
Mutations in this gene have been associated with idiopathic cardiomyopathy (IDC), premature aging, sporadic motor neuron disease. The dysfunction of SOD2 may lead to increased intracellular oxidative stress, which in turn is involved in the occurrence and development of some neurodegenerative diseases (such as Alzheimer's disease, Parkinson's disease), nervous system damage, and lung diseases. In addition, some studies have found that the expression level of SOD2 is associated with tumorigenesis and treatment response.
Bioapplications of SOD2
SOD2 protein has many applications in the field of biology, involving cellular oxidative stress, mitochondrial function, neurodegenerative diseases, tumorigenesis and development. The in-depth study of it is helpful to reveal the pathogenesis of related diseases and provide new targets and strategies for the treatment of diseases.
Case Study
Case study 1: Diego R Coelho, 2022
The dichotomous behavior of superoxide dismutase-2 (SOD2) in cancer biology has long been acknowledged and more recently linked to different posttranslational forms of the enzyme. However, a distinctive activity underlying its tumor-promoting function is yet to be described. Western blots confirming the subcellular localization of SOD2 were performed and a PicoProbe Formaldehyde Assay Kit (BioVision, K805) was used to quantify formaldehyde (HCHO) produced by demethylase activity of SOD2 and so on.
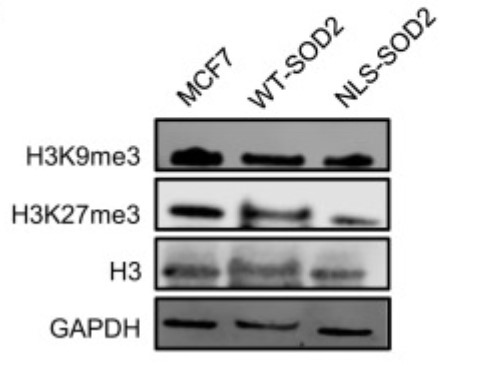
Here, the researchers report that acetylation, one of such posttranslational modifications (PTMs), increases SOD2 affinity for iron, effectively changing the biochemical function of this enzyme from that of an antioxidant to a demethylase. Acetylated, iron-bound SOD2 localizes to the nucleus, promoting stem cell gene expression via removal of suppressive epigenetic marks such as H3K9me3 and H3K927me3. Together, the results provide evidence in support of an acetylation-dependent nucleus-specific function of SOD2 associated with the emergence of more tumorigenic and metastatic cancer cell phenotypes.
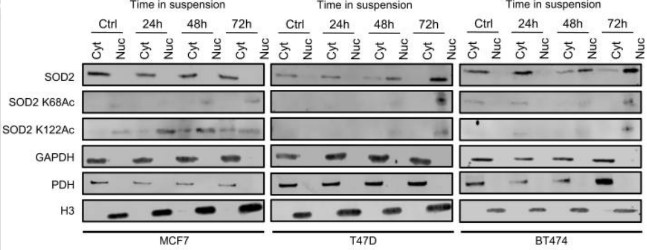
Fig1. Acetylated SOD2 accumulates in the nuclear fraction of breast cancer cell lines cultured in suspension.
Case study 2: Atinuke Dosunmu-Ogunbi, 2022
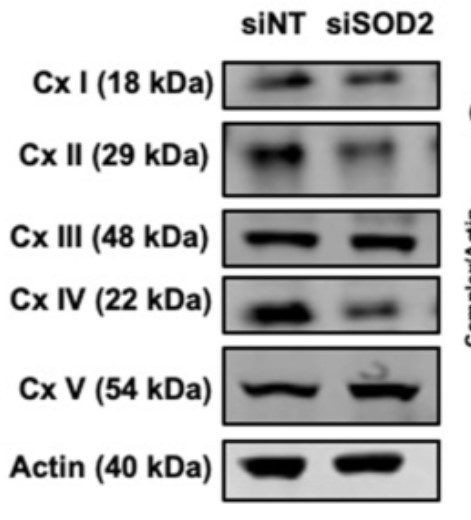
Sickle cell disease (SCD) is a genetic red blood cell disorder characterized by increased reactive oxygen species (ROS) and a concordant reduction in antioxidant capacity in the endothelium. Superoxide dismutase 2 (SOD2) is a mitochondrial-localized enzyme that catalyzes the dismutation of superoxide to hydrogen peroxide. Decreased peripheral blood expression of SOD2 is correlated with increased hemolysis and cardiomyopathy in SCD.
To investigate the impact of decreased SOD2 expression in the endothelium, SOD2 was knocked down in human pulmonary microvascular endothelial cells (hPMVECs). The researchers found that SOD2 deficiency in hPMVECs results in endothelial cell dysfunction, including reduced cellular adhesion, diminished migration, integrin protein dysregulation, and disruption of permeability. Furthermore, they uncover that SOD2's inability to facilitate dimerization. These results demonstrate that endothelial cells are deficient in SOD2 expression in SCD patients and suggest a novel pathway for SOD2 in regulating fibronectin processing.
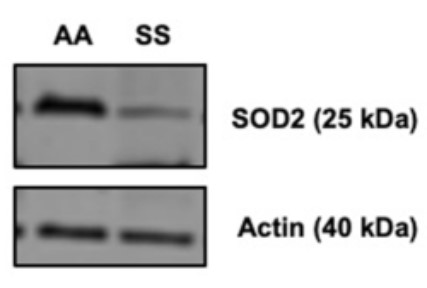
Fig3. Townes transgenic SS mice compared to AA mice have decreased SOD2 protein expression in whole lung lysate.
Quality Guarantee
High Purity
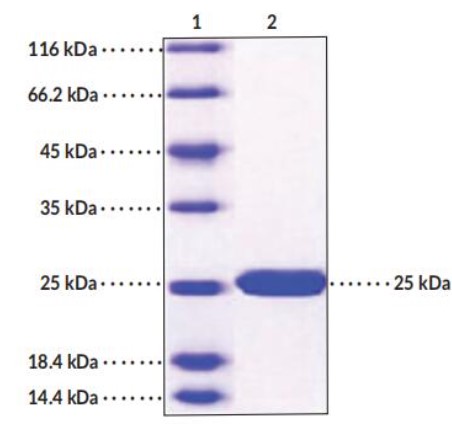
Fig1. SDS-PAGE (SOD2-06H) (PROTOCOL for western blot)
.
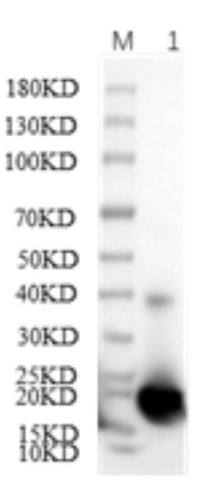
Fig2. SDS-PAGE (SOD2-02H) (PROTOCOL for western blot)
Involved Pathway
SOD2 involved in several pathways and played different roles in them. We selected most pathways SOD2 participated on our site, such as FoxO signaling pathway,Peroxisome,Huntingtons disease, which may be useful for your reference. Also, other proteins which involved in the same pathway with SOD2 were listed below. Creative BioMart supplied nearly all the proteins listed, you can search them on our site.
| Pathway Name | Pathway Related Protein |
|---|---|
| Peroxisome | PEX5L,ACOX1,AMACR,ACSL3B,CAT,FAR2,NOS2,PIPOX,CRATA,HMGCLL1 |
| FoxO signaling pathway | PRKAG2,TGFBR1A,BRAF,PRKAG3B,STAT3,TGFBR1B,SMAD4,IL6,PTEN,CDKN2D |
| Huntingtons disease | TGM2,NDUFS8,UCP1,NDUFA11,TP53,NDUFB1,GRM5,DNAH14,NDUFV3,COX7A2 |
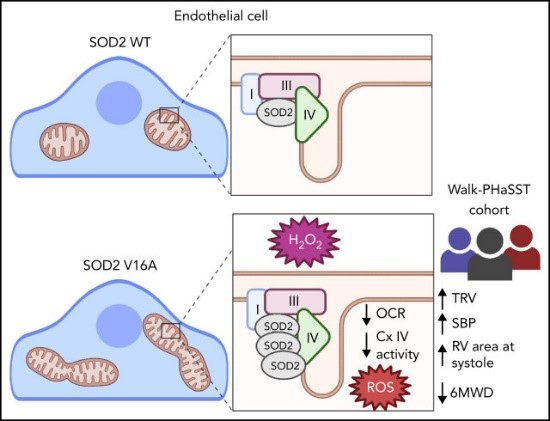
Fig1. SOD2V16A is a novel clinical biomarker of cardiovascular dysfunction in SCD patients through its ability to decrease mitochondrial complex IV activity and amplify ROS production in the endothelium. (Atinuke Dosunmu-Ogunbi, 2022)
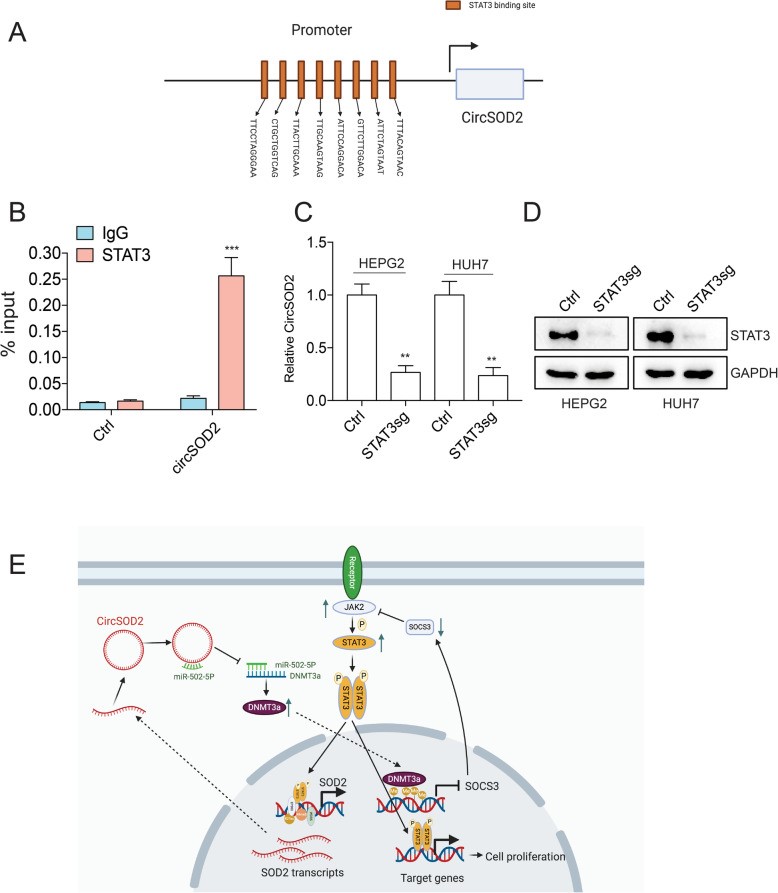
Fig2. The novel signaling axis circSOD2/miR-502-5p/DNMT3a/JAK2/STAT3/circSOD2 provides a better understanding of HCC tumorigenesis. (Zhongwei Zhao, 2020)
Protein Function
SOD2 has several biochemical functions, for example, identical protein binding,manganese ion binding,superoxide dismutase activity. Some of the functions are cooperated with other proteins, some of the functions could acted by SOD2 itself. We selected most functions SOD2 had, and list some proteins which have the same functions with SOD2. You can find most of the proteins on our site.
| Function | Related Protein |
|---|---|
| identical protein binding | LDHA,CD28,TMEM115,MBIP,SH3BP4,MRFAP1L1,USP4,CAD,GOT2A,VEGFA |
| manganese ion binding | B4GALT1,GLUL,MPPE1,GALNT2,TDP2B,ME1,PPM1M,CDIPT,PPM1AA,NPEPL1 |
| superoxide dismutase activity | SOD3A,PRNPA,NQO1,CCS,SOD1 |
Interacting Protein
SOD2 has direct interactions with proteins and molecules. Those interactions were detected by several methods such as yeast two hybrid, co-IP, pull-down and so on. We selected proteins and molecules interacted with SOD2 here. Most of them are supplied by our site. Hope this information will be useful for your research of SOD2.
AURKA;RIPK2;SMARCA5;EIF1B;CST4;CST1;AMY1B;P4HB;MYH8;HSP90B1;MCC;MPG;HLA-B;VHL;RPL21;SMARCA2;ARF6;SCRN1
Resources
Research Area
Related Services
Related Products
References
- Kang, PT; Chen, CL; et al. BCNU-induced gR2 DEFECT mediates S-glutathionylation of Complex I and respiratory uncoupling in myocardium. BIOCHEMICAL PHARMACOLOGY 89:490-502(2014).
- Kang, PT; Chen, CL; et al. Increased mitochondrial prooxidant activity mediates up-regulation of Complex I S-glutathionylation via protein thiyl radical in the murine heart of eNOS(-/-). FREE RADICAL BIOLOGY AND MEDICINE 79:56-68(2015).



