Oncoprotein-Signal Transducers
Related Symbol Search List
- ABL1
- AKT1
- AKT2
- BRAF
- CRK
- CTNNB1
- FES
- FGR
- FLT3
- GIP
- GNAS
- HRAS
- JAK1
- JAK2
- KRAS
- MCF2
- MOS
- MYD88
- NRAS
- PIK3CA
- PIM1
- RAF1
- Src
- VAV1
- YES1
Immunology Background
 Figure 1. Signal transduction signal pathway protein gene.
Figure 1. Signal transduction signal pathway protein gene.
In the signal transduction system, there are many important proteins, such as G-protein, Ras protein plays the role of molecular switch, CKI1 and GCR1 may act as cytokinin receptors and participate in cytokinin signal transduction. Scaffold proteins are proteins that do not have enzyme activity in themselves, acting like molecular glue, bonding function-related proteins together, and so on. These proteins play an important role in signal transduction. Please follow me to learn more about these proteins.
What is cellular signal transduction?
Cell signal transduction refers to the process by which extracellular factors bind to receptors, triggering a series of biochemical reactions and protein-protein interactions in the cell, until the genes required for the physiological response of the cell begin to be expressed and various biological effects are formed. It is now known that there are multiple signal transduction methods and pathways in cells, which is a very complicated network system. In a cell communication system, cells either identify the cells that are in contact with them, or recognize the various signals present in the surrounding environment, and convert them into changes in the function of various molecules within the cell, thereby changing certain metabolisms within the cell Process, affect the growth rate of cells, and even induce cell death. This change in the activity of various molecules that occurs against exogenous signals, and the process of transmitting this change to effector molecules in order to change the function of cells is called signal transduction, and its ultimate purpose is to make the body The most appropriate response is to changes in the external environment. The regulation of substance metabolism often involves the regulation of the metabolic pathways at the overall level by the neuro-endocrine system. The essence is that some cells in the body send out signals, and another part of the cells receives the signals and transforms them into changes in cell function. Therefore, to clarify the mechanism of cell signal transduction means to understand the cell's performance and regulation in the aspects of proliferation, differentiation, metabolism and death in the whole life process, and then to understand the regulation mechanism of body growth, development and metabolism.
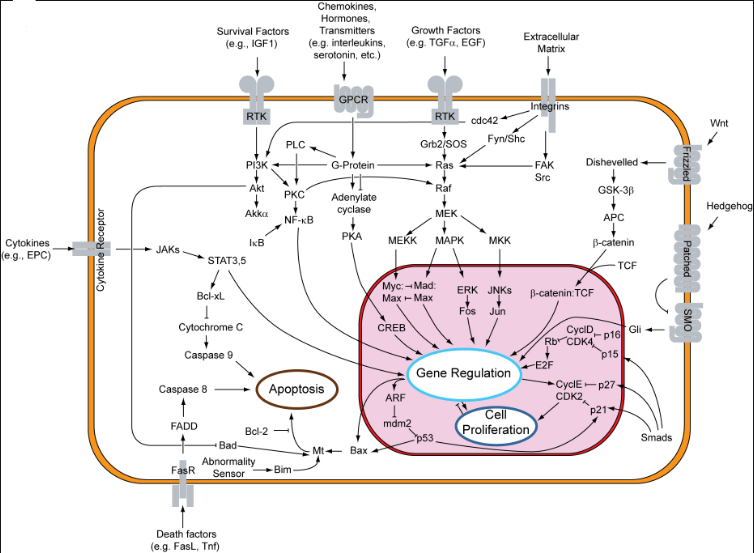 Figure 2. Cell proliferation, apoptosis-related gene expression and signal transduction pathway. (By Shi Xiuchao Science Network blog.)
Figure 2. Cell proliferation, apoptosis-related gene expression and signal transduction pathway. (By Shi Xiuchao Science Network blog.)
Signal transducers Proteins
There are many kinds of signal transduction proteins, which can be divided into signal transduction proteins related to molecular switches, proteins involved in cytokinin signal transduction, junction proteins and tendril proteins.
1. Signal transduction proteins related to molecular switches
(1)G-protein is a kind of signal transduction protein which can bind to guanosine diphosphate and has the activity of GTP hydrolase. When a cell transduces extracellular signals, various extracellular ligands are first received by different types of G-protein-coupled receptors, and then the receptors are activated to further activate the heterotrimeric G-protein on the inside of the plasma membrane, which then activates various effectors downstream to produce a second messenger in the cell. Thus, the signal is transmitted step by step to regulate the growth and development process of the organism. GTP-bindinG-protein, which plays an important role in intracellular signal transduction pathway, is composed of three different subunits: α, β and γ. Hormone binds to hormone receptor and induces GTP to exchange GDP bound with G-protein. Activated G-protein can activate adenylate cyclase in the middle and lower reaches of signal transduction pathway. G-protein connects the extracellular first messenger epinephrine and the second signal of adenylate cyclase-catalyzed adenylate cyclase to cAMP.
(2)Ras protein is a key component of the RTK-mediated signaling pathway. When the ligand binds to the receptor, activating the tyrosine protein kinase activity of the receptor causes a series of phosphorylation cascades, resulting in changes in cell physiology or gene expression. This signal pathway completes the signal transduction through the activation of tyrosine protein kinase of the receptor itself. Ras protein is the expression product of Ras gene. Ras protein has GTPase activity, which is activated when binding to GTP and inactivated when binding to GDP, so Ras protein has the function of molecular switch.
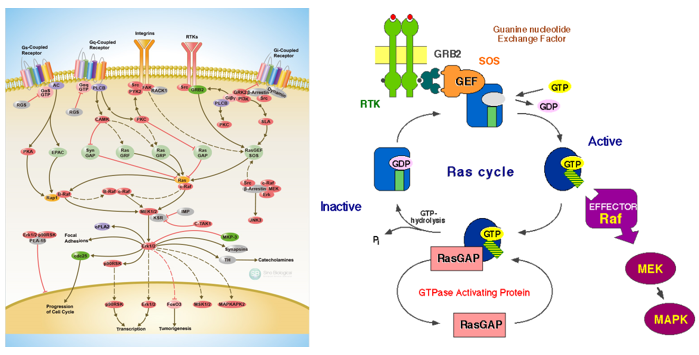 Figure 3. G protein associated receptor signal pathway. (By sinobiological.) G protein Ras signal pathway map. (By bioon. )
Figure 3. G protein associated receptor signal pathway. (By sinobiological.) G protein Ras signal pathway map. (By bioon. )
2. Proteins involved in cytokinin signal transduction
CKI1 protein. The main elements of the bacterial binary regulatory system are histidine kinase (infectious protein) and response regulatory protein (receptive protein), while the products encoded by CKI1 gene have regulatory protein regions and sequences similar to histidine kinases, and highly conserved amino acid residues in histidine kinases, including automatic phosphorylation site His405, are found in CKI1. In addition, CKI1 protein has a unique N-terminal, which includes two hypothetical transmembrane domains, and its extracellular domain may serve as a cytokinin binding site. When CKI1 protein is overexpressed, it causes typical cytokinin response.
3. Junction proteins
The junction protein c-Crk is the product of the expression of proto-oncogene c-Crk, which participates in the signal transduction of many molecules such as receptor tyrosine kinase and integrin. The SH2 and SH3 domains in Crk play an important role in the realization of protein function. The role of Crk in receptor tyrosine kinase signal transduction. Under the stimulation of insulin or insulin-like growth factor-1, tyrosine phosphorylation of c-CrkⅡ occurs, which is the first evidence that Crk is involved in insulin receptor signal. Further studies showed that c-CrkⅡ could bind to islet receptors. In CHO cells, insulin induced phosphorylation of c-CrkⅡ and dissociation of Crk-C3G complex. This dissociation caused conformational changes of Rap1, a downstream molecule of C3G, and affected the enzyme activity of Rap1. C3G is the main downstream signal molecule of Crk. It is a guanosine monophosphate releasing factor of small G protein Rap1, which can transform Rap1 from Rap1-GDP to active form Rap1-GTP. Rap1 can inhibit M-APK system by competitively binding to Raf1.
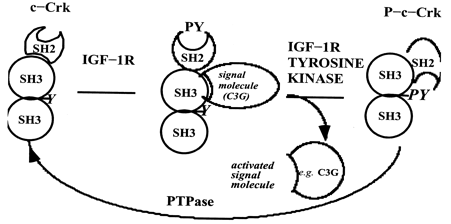 Figure 4. Model illustrating the role of c-Crk in the IGF-1-initiated component of induction of adipocyte differentiation. (Jin S, et al. 2000 )
Figure 4. Model illustrating the role of c-Crk in the IGF-1-initiated component of induction of adipocyte differentiation. (Jin S, et al. 2000 )
4.Tendril proteins
At present, it is considered that the biochemical function of CLV1 is an upstream event of RLKs involved in signal transduction. The signal cascade is amplified by protein interaction to regulate the meristem. The molecular weight of CLV1 1 is close to 105 ku, and it is easy to be degraded. The interaction between CLV1 and KAPP suggests that the phosphorylation cascade system depends on the automatic phosphorylation of CLV1 to some extent.As the main manufacturer of recombinant protein, Creative Biomart provides Cell survival & death Proteins, from a variety of sources, grades and formulations to facilitate your efficient and accurate research content in the experiment.
Reference:
- Jin S, Zhai B, Qiu Z, et al. c-Crk, a substrate of the insulin-like growth factor-1 receptor tyrosine kinase, functions as an early signal mediator in the adipocyte differentiation process. Journal of Biological Chemistry, 2000, 275(44): 34344-34352.

